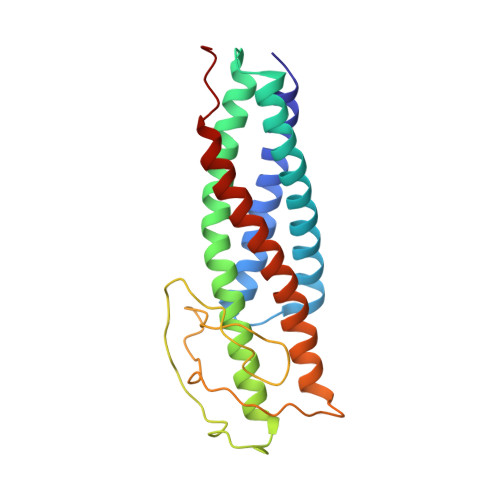
Chaperone Recycling in Late-Stage Flagellar Assembly.
Rossi, P., Xing, Q., Bini, E., Portaliou, A.G., Clay, M.C., Warren, E.M., Khanra, N.K., Economou, A., Kalodimos, C.G.(2023) J Mol Biology 435: 167954-167954
- PubMed: 37330284
- DOI: https://doi.org/10.1016/j.jmb.2023.167954
- Primary Citation Related Structures:
8FTW, 8FTX - PubMed Abstract:
The flagellum is a sophisticated nanomachine responsible for motility in Gram-negative bacteria. Flagellar assembly is a strictly choreographed process, in which the motor and export gate are formed first, followed by the extracellular propeller structure. Extracellular flagellar components are escorted to the export gate by dedicated molecular chaperones for secretion and self-assembly at the apex of the emerging structure. The detailed mechanisms of chaperone-substrate trafficking at the export gate remain poorly understood. Here, we structurally characterized the interaction of Salmonella enterica late-stage flagellar chaperones FliT and FlgN with the export controller protein FliJ. Previous studies showed that FliJ is absolutely required for flagellar assembly since its interaction with chaperone-client complexes controls substrate delivery to the export gate. Our biophysical and cell-based data show that FliT and FlgN bind FliJ cooperatively, with high affinity and on specific sites. Chaperone binding completely disrupts the FliJ coiled-coil structure and alters its interactions with the export gate. We propose that FliJ aids the release of substrates from the chaperone and forms the basis of chaperone recycling during late-stage flagellar assembly.
- Deparment of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN 38105, United States.
Organizational Affiliation: