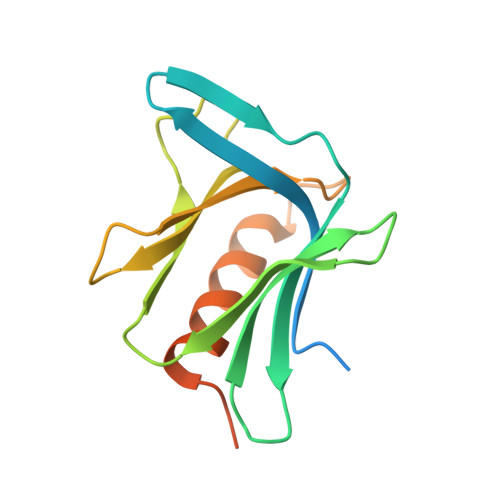
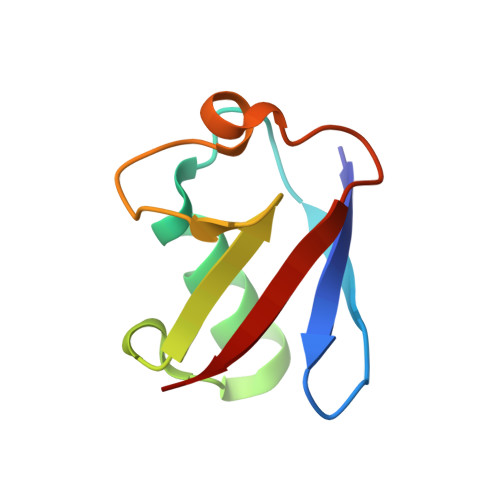
A structure-based designed small molecule depletes hRpn13 Pru and a select group of KEN box proteins.
Lu, X., Chandravanshi, M., Sabbasani, V.R., Gaikwad, S., Hughitt, V.K., Gyabaah-Kessie, N., Scroggins, B.T., Das, S., Myint, W., Clapp, M.E., Schwieters, C.D., Dyba, M.A., Bolhuis, D.L., Koscielniak, J.W., Andresson, T., Emanuele, M.J., Brown, N.G., Matsuo, H., Chari, R., Citrin, D.E., Mock, B.A., Swenson, R.E., Walters, K.J.(2024) Nat Commun 15: 2485-2485
- PubMed: 38509117 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46644-7
- Primary Citation Related Structures:
8FTQ - PubMed Abstract:
Proteasome subunit hRpn13 is partially proteolyzed in certain cancer cell types to generate hRpn13 Pru by degradation of its UCHL5/Uch37-binding DEUBAD domain and retention of an intact proteasome- and ubiquitin-binding Pru domain. By using structure-guided virtual screening, we identify an hRpn13 binder (XL44) and solve its structure ligated to hRpn13 Pru by integrated X-ray crystallography and NMR to reveal its targeting mechanism. Surprisingly, hRpn13 Pru is depleted in myeloma cells following treatment with XL44. TMT-MS experiments reveal a select group of off-targets, including PCNA clamp-associated factor PCLAF and ribonucleoside-diphosphate reductase subunit M2 (RRM2), that are similarly depleted by XL44 treatment. XL44 induces hRpn13-dependent apoptosis and also restricts cell viability by a PCLAF-dependent mechanism. A KEN box, but not ubiquitination, is required for XL44-induced depletion of PCLAF. Here, we show that XL44 induces ubiquitin-dependent loss of hRpn13 Pru and ubiquitin-independent loss of select KEN box containing proteins.
- Protein Processing Section, Center for Structural Biology, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Frederick, MD, USA.
Organizational Affiliation: