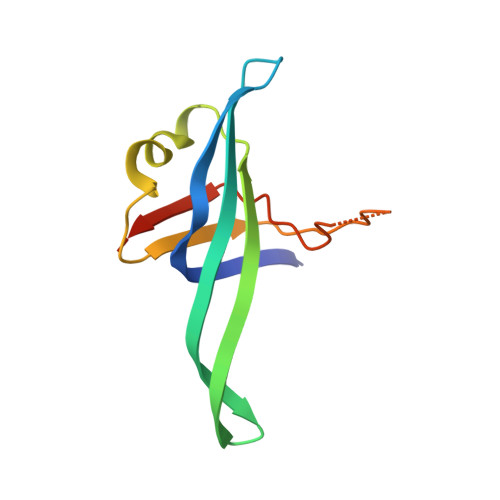
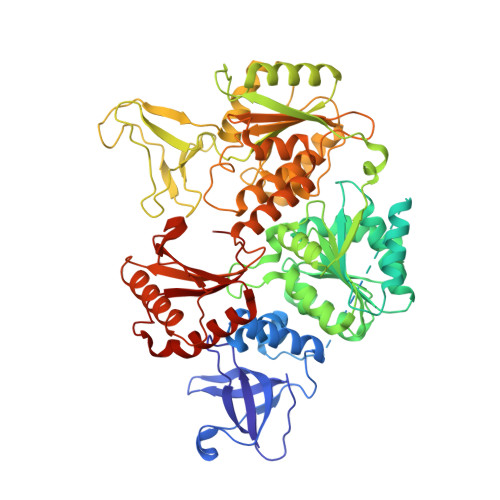
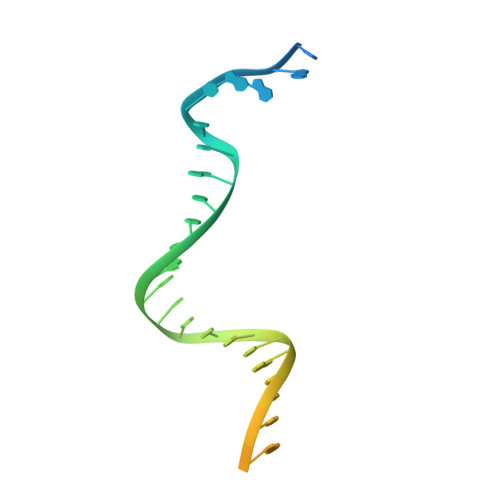
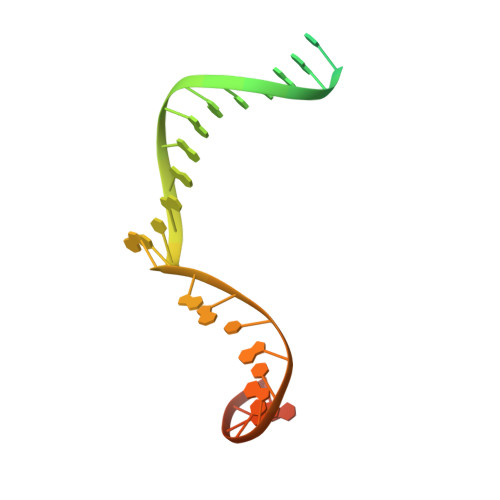
Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli.
Duckworth, A.T., Ducos, P.L., McMillan, S.D., Satyshur, K.A., Blumenthal, K.H., Deorio, H.R., Larson, J.A., Sandler, S.J., Grant, T., Keck, J.L.(2023) Nat Commun 14: 2725-2725
- PubMed: 37169801 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38144-x
- Primary Citation Related Structures:
8FAK - PubMed Abstract:
Bacterial replisomes often dissociate from replication forks before chromosomal replication is complete. To avoid the lethal consequences of such situations, bacteria have evolved replication restart pathways that reload replisomes onto prematurely terminated replication forks. To understand how the primary replication restart pathway in E. coli (PriA-PriB) selectively acts on replication forks, we determined the cryogenic-electron microscopy structure of a PriA/PriB/replication fork complex. Replication fork specificity arises from extensive PriA interactions with each arm of the branched DNA. These interactions reshape the PriA protein to create a pore encircling single-stranded lagging-strand DNA while also exposing a surface of PriA onto which PriB docks. Together with supporting biochemical and genetic studies, the structure reveals a switch-like mechanism for replication restart initiation in which restructuring of PriA directly couples replication fork recognition to PriA/PriB complex formation to ensure robust and high-fidelity replication re-initiation.
- Department of Biomolecular Chemistry, University of Wisconsin-Madison, Madison, WI, 53706, USA.
Organizational Affiliation: