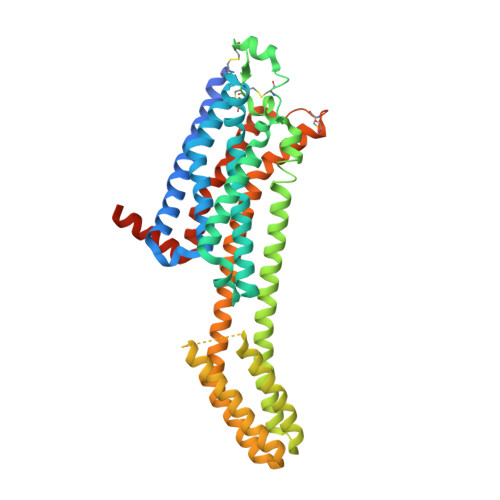
High ligand efficiency quinazoline compounds as novel A 2A adenosine receptor antagonists.
Bolteau, R., Duroux, R., Laversin, A., Vreulz, B., Shiriaeva, A., Stauch, B., Han, G.W., Cherezov, V., Renault, N., Barczyk, A., Ravez, S., Coevoet, M., Melnyk, P., Liberelle, M., Yous, S.(2022) Eur J Med Chem 241: 114620-114620
- PubMed: 35933788 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2022.114620
- Primary Citation Related Structures:
8DU3 - PubMed Abstract:
The past fifty years have been marked by the surge of neurodegenerative diseases. Unfortunately, current treatments are only symptomatic. Hence, the search for new and innovative therapeutic targets for curative treatments becomes a major challenge. Among these targets, the adenosine A 2A receptor (A 2A AR) has been the subject of much research in recent years. In this paper, we report the design, synthesis and pharmacological analysis of quinazoline derivatives as A 2A AR antagonists with high ligand efficiency. This class of molecules has been discovered by a virtual screening and bears no structural semblance with reference antagonist ZM-241385. More precisely, we identified a series of 2-aminoquinazoline as promising A 2A AR antagonists. Among them, one compound showed a high affinity towards A 2A AR (21a, K i = 20 nM). We crystallized this ligand in complex with A 2A AR, confirming one of our predicted docking poses and opening up possibilities for further optimization to derive selective ligands for specific adenosine receptor subtypes.
- Univ. Lille, Inserm, CHU Lille, U1172, LilNCog - Lille Neuroscience & Cognition, F-59000, Lille, France.
Organizational Affiliation: