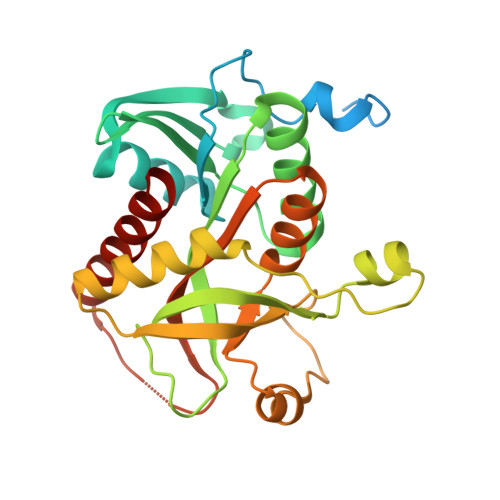
The structure of His-tagged Geobacillus stearothermophilus purine nucleoside phosphorylase reveals a 'spanner in the works'.
Given, F.M., Moran, F., Johns, A.S., Titterington, J.A., Allison, T.M., Crittenden, D.L., Johnston, J.M.(2022) Acta Crystallogr F Struct Biol Commun 78: 416-422
- PubMed: 36458621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22011025
- Primary Citation Related Structures:
8D38 - PubMed Abstract:
The 1.72 Å resolution structure of purine nucleoside phosphorylase from Geobacillus stearothermophilus, a thermostable protein of potential interest for the biocatalytic synthesis of antiviral nucleoside compounds, is reported. The structure of the N-terminally His-tagged enzyme is a hexamer, as is typical of bacterial homologues, with a trimer-of-dimers arrangement. Unexpectedly, several residues of the recombinant tobacco etch virus protease (rTEV) cleavage site from the N-terminal tag are located in the active site of the neighbouring subunit in the dimer. Key to this interaction is a tyrosine residue, which sits where the nucleoside ring of the substrate would normally be located. Tag binding appears to be driven by a combination of enthalpic, entropic and proximity effects, which convey a particularly high affinity in the crystallized form. Attempts to cleave the tag in solution yielded only a small fraction of untagged protein, suggesting that the enzyme predominantly exists in the tag-bound form in solution, preventing rTEV from accessing the cleavage site. However, the tagged protein retained some activity in solution, suggesting that the tag does not completely block the active site, but may act as a competitive inhibitor. This serves as a warning that it is prudent to establish how affinity tags may affect protein structure and function, especially for industrial biocatalytic applications that rely on the efficiency and convenience of one-pot purifications and in cases where tag removal is difficult.
- School of Physical and Chemical Sciences, Biomolecular Interaction Centre, University of Canterbury, New Zealand.
Organizational Affiliation: