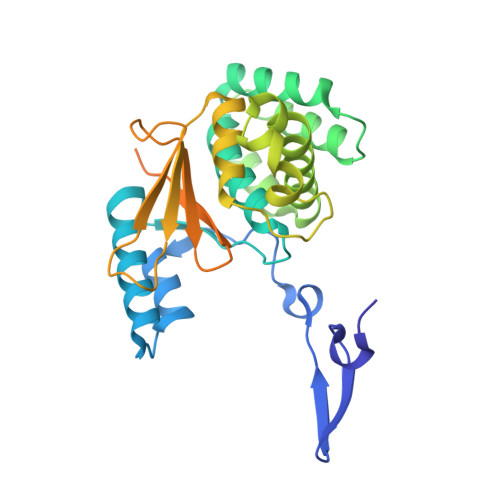
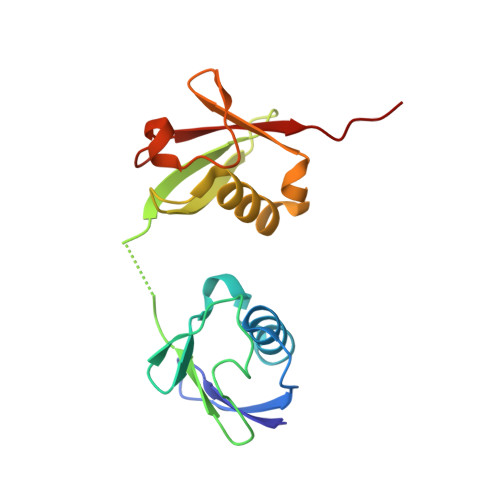
Functional and structural diversity in deubiquitinases of the Chlamydia-like bacterium Simkania negevensis.
Boll, V., Hermanns, T., Uthoff, M., Erven, I., Horner, E.M., Kozjak-Pavlovic, V., Baumann, U., Hofmann, K.(2023) Nat Commun 14: 7335-7335
- PubMed: 37957213 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-43144-y
- Primary Citation Related Structures:
8CMR - PubMed Abstract:
Besides the regulation of many cellular pathways, ubiquitination is important for defense against invading pathogens. Some intracellular bacteria have evolved deubiquitinase (DUB) effector proteins, which interfere with the host ubiquitin system and help the pathogen to evade xenophagy and lysosomal degradation. Most intracellular bacteria encode one or two DUBs, which are often linkage-promiscuous or preferentially cleave K63-linked chains attached to bacteria or bacteria-containing vacuoles. By contrast, the respiratory pathogen Legionella pneumophila possesses a much larger number of DUB effectors, including a K6-specific enzyme belonging to the OTU family and an M1-specific DUB uniquely found in this bacterium. Here, we report that the opportunistic pathogen Simkania negevensis, which is unrelated to Legionella but has a similar lifestyle, encodes a similarly large number of DUBs, including M1- and K6-specific enzymes. Simkania DUBs are highly diverse and include DUB classes never before seen in bacteria. Interestingly, the M1- and K6-specific DUBs of Legionella and Simkania are unrelated, suggesting that their acquisition occurred independently. We characterize the DUB activity of eight Simkania-encoded enzymes belonging to five different DUB classes. We also provide a structural basis for the M1-specificity of a Simkania DUB, which most likely evolved from a eukaryotic otubain-like precursor.
- Institute for Genetics, University of Cologne, Cologne, Germany.
Organizational Affiliation: