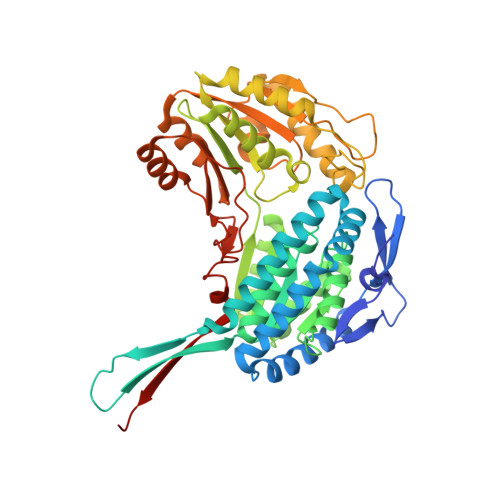
Molecular basis of sulfolactate synthesis by sulfolactaldehyde dehydrogenase from Rhizobium leguminosarum.
Li, J., Sharma, M., Meek, R., Alhifthi, A., Armstrong, Z., Soler, N.M., Lee, M., Goddard-Borger, E.D., Blaza, J.N., Davies, G.J., Williams, S.J.(2023) Chem Sci 14: 11429-11440
- PubMed: 37886098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d3sc01594g
- Primary Citation Related Structures:
8C54 - PubMed Abstract:
Sulfolactate (SL) is a short-chain organosulfonate that is an important reservoir of sulfur in the biosphere. SL is produced by oxidation of sulfolactaldehyde (SLA), which in turn derives from sulfoglycolysis of the sulfosugar sulfoquinovose, or through oxidation of 2,3-dihydroxypropanesulfonate. Oxidation of SLA is catalyzed by SLA dehydrogenases belonging to the aldehyde dehydrogenase superfamily. We report that SLA dehydrogenase Rl GabD from the sulfoglycolytic bacterium Rhizobium leguminsarum SRDI565 can use both NAD + and NADP + as cofactor to oxidize SLA, and indicatively operates through a rapid equilibrium ordered mechanism. We report the cryo-EM structure of Rl GabD bound to NADH, revealing a tetrameric quaternary structure and supporting proposal of organosulfonate binding residues in the active site, and a catalytic mechanism. Sequence based homology searches identified SLA dehydrogenase homologs in a range of putative sulfoglycolytic gene clusters in bacteria predominantly from the phyla Actinobacteria, Firmicutes, and Proteobacteria. This work provides a structural and biochemical view of SLA dehydrogenases to complement our knowledge of SLA reductases, and provide detailed insights into a critical step in the organosulfur cycle.
- School of Chemistry and Bio21 Molecular Science and Biotechnology Institute, University of Melbourne Parkville Victoria 3010 Australia sjwill@unimelb.edu.au.
Organizational Affiliation: