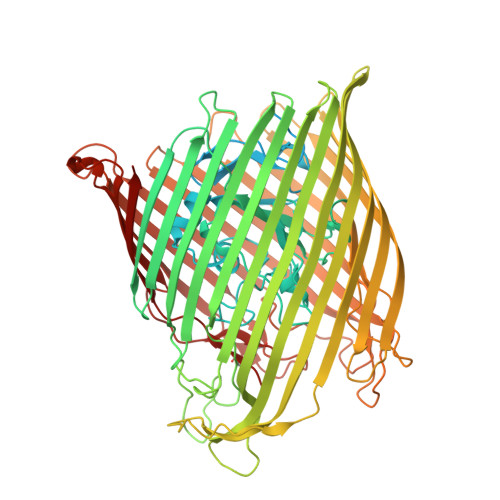
Interactions of TonB-dependent transporter FoxA with siderophores and antibiotics that affect binding, uptake, and signal transduction.
Chan, D.C.K., Josts, I., Koteva, K., Wright, G.D., Tidow, H., Burrows, L.L.(2023) Proc Natl Acad Sci U S A 120: e2221253120-e2221253120
- PubMed: 37043535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2221253120
- Primary Citation Related Structures:
8B43 - PubMed Abstract:
The outer membrane of gram-negative bacteria prevents many antibiotics from reaching intracellular targets. However, some antimicrobials can take advantage of iron import transporters to cross this barrier. We showed previously that the thiopeptide antibiotic thiocillin exploits the nocardamine xenosiderophore transporter, FoxA, of the opportunistic pathogen Pseudomonas aeruginosa for uptake. Here, we show that FoxA also transports the xenosiderophore bisucaberin and describe at 2.5 Å resolution the crystal structure of bisucaberin bound to FoxA. Bisucaberin is distinct from other siderophores because it forms a 3:2 rather than 1:1 siderophore-iron complex. Mutations in a single extracellular loop of FoxA differentially affected nocardamine, thiocillin, and bisucaberin binding, uptake, and signal transduction. These results show that in addition to modulating ligand binding, the extracellular loops of siderophore transporters are of fundamental importance for controlling ligand uptake and its regulatory consequences, which have implications for the development of siderophore-antibiotic conjugates to treat difficult infections.
- David Braley Center for Antibiotic Discovery, McMaster University, Hamilton, ON L8S 4K1, Canada.
Organizational Affiliation: