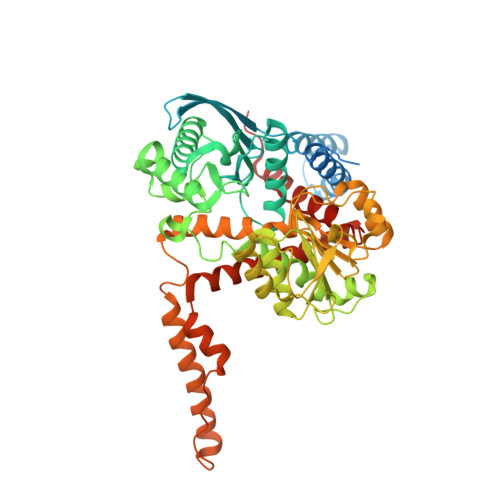
Structural Basis for the Binding of Allosteric Activators Leucine and ADP to Mammalian Glutamate Dehydrogenase.
Aleshin, V.A., Bunik, V.I., Bruch, E.M., Bellinzoni, M.(2022) Int J Mol Sci 23
- PubMed: 36232607
- DOI: https://doi.org/10.3390/ijms231911306
- Primary Citation of Related Structures:
8AR7, 8AR8 - PubMed Abstract:
Glutamate dehydrogenase (GDH) plays a key role in the metabolism of glutamate, an important compound at a cross-road of carbon and nitrogen metabolism and a relevant neurotransmitter. Despite being one of the first discovered allosteric enzymes, GDH still poses challenges for structural characterization of its allosteric sites. Only the structures with ADP, and at low (3.5 Å) resolution, are available for mammalian GDH complexes with allosteric activators. Here, we aim at deciphering a structural basis for the GDH allosteric activation using bovine GDH as a model. For the first time, we report a mammalian GDH structure in a ternary complex with the activators leucine and ADP, co-crystallized with potassium ion, resolved to 2.45 Å. An improved 2.4-angstrom resolution of the GDH complex with ADP is also presented. The ternary complex with leucine and ADP differs from the binary complex with ADP by the conformation of GDH C-terminus, involved in the leucine binding and subunit interactions. The potassium site, identified in this work, may mediate interactions between the leucine and ADP binding sites. Our data provide novel insights into the mechanisms of GDH activation by leucine and ADP, linked to the enzyme regulation by (de)acetylation.
- Department of Biokinetics, A. N. Belozersky Institute of Physicochemical Biology, Lomonosov Moscow State University, 119234 Moscow, Russia.
Organizational Affiliation: