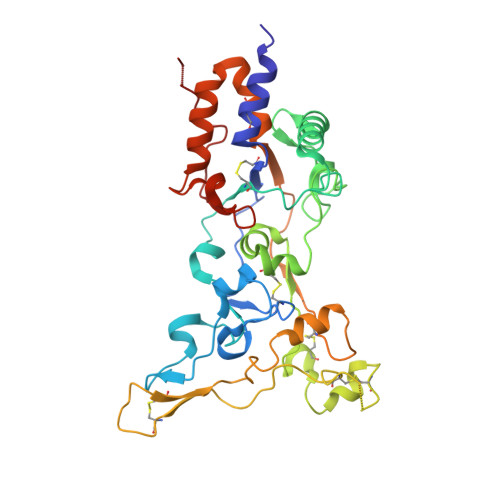
The crystal structure of a simian Foamy Virus receptor binding domain provides clues about entry into host cells.
Fernandez, I., Dynesen, L.T., Coquin, Y., Pederzoli, R., Brun, D., Haouz, A., Gessain, A., Rey, F.A., Buseyne, F., Backovic, M.(2023) Nat Commun 14: 1262-1262
- PubMed: 36878926 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36923-0
- Primary Citation Related Structures:
8AEZ, 8AIC - PubMed Abstract:
The surface envelope glycoprotein (Env) of all retroviruses mediates virus binding to cells and fusion of the viral and cellular membranes. A structure-function relationship for the HIV Env that belongs to the Orthoretrovirus subfamily has been well established. Structural information is however largely missing for the Env of Foamy viruses (FVs), the second retroviral subfamily. In this work we present the X-ray structure of the receptor binding domain (RBD) of a simian FV Env at 2.57 Å resolution, revealing two subdomains and an unprecedented fold. We have generated a model for the organization of the RBDs within the trimeric Env, which indicates that the upper subdomains form a cage-like structure at the apex of the Env, and identified residues K342, R343, R359 and R369 in the lower subdomain as key players for the interaction of the RBD and viral particles with heparan sulfate.
- Institut Pasteur, Université Paris Cité, CNRS UMR3569, Unité de Virologie Structurale, 75015, Paris, France.
Organizational Affiliation: