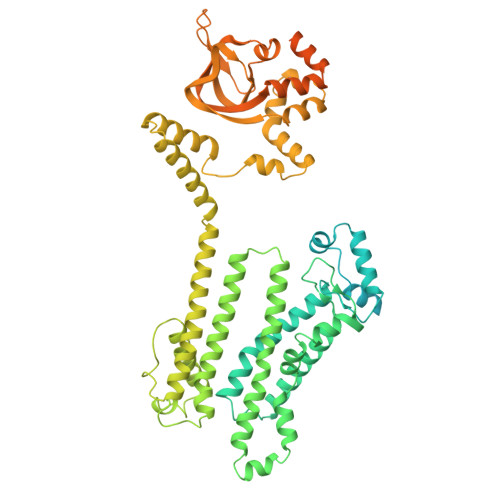
The HCN Channel Voltage Sensor Undergoes A Large Downward Motion During Hyperpolarization
Dai, G., Aman, T.K., DiMaio, F., Zagotta, W.N.(2019) Nat Struct Mol Biol 26: 686-694
- PubMed: 31285608
- DOI: https://doi.org/10.1038/s41594-019-0259-1
- Primary Citation of Related Structures:
8ZZW - PubMed Abstract:
Voltage-gated ion channels (VGICs) contain positively charged residues within the S4 helix of the voltage-sensing domain (VSD) that are displaced in response to changes in transmembrane voltage, promoting conformational changes that open the pore. Pacemaker hyperpolarization-activated cyclic nucleotide-gated (HCN) channels are unique among VGICs because their open probability is increased by membrane hyperpolarization rather than depolarization. Here we measured the precise movement of the S4 helix of a sea urchin HCN channel using transition metal ion fluorescence resonance energy transfer (tmFRET). We show that the S4 undergoes a substantial (~10 Å) downward movement in response to membrane hyperpolarization. Furthermore, by applying distance constraints determined from tmFRET experiments to Rosetta modeling, we reveal that the carboxy-terminal part of the S4 helix exhibits an unexpected tilting motion during hyperpolarization activation. These data provide a long-sought glimpse of the hyperpolarized state of a functioning VSD and also a framework for understanding the dynamics of reverse gating in HCN channels.
- Department of Physiology and Biophysics, University of Washington, Seattle, WA, USA.
Organizational Affiliation: