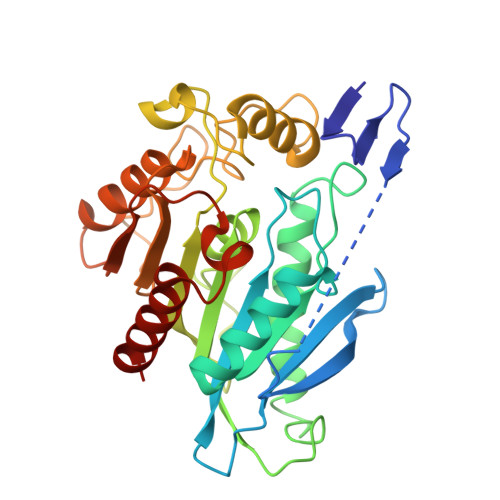
Molecular Basis for Catalysis and Regulation of the Strigolactone Catabolic Enzyme CXE15.
Shahul Hameed, U.F., Balakrishna, A., Wang, J.Y., Alvarez, D., Momin, A.A., Schwarzenberg, A., Al-Babili, S., Arold, S.T.(2025) Nat Commun 16: 10290-10290
- PubMed: 41271671 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-65204-1
- Primary Citation Related Structures:
8ZR6, 8ZRF, 8ZRG, 8ZRO - PubMed Abstract:
Strigolactones (SLs) are pivotal plant hormones involved in developmental, physiological, and adaptive processes. SLs also facilitate symbiosis with arbuscular mycorrhizal fungi and trigger germination of root parasitic Striga plants. The carboxylesterase CXE15, recently identified as the SL catabolic enzyme in Arabidopsis thaliana, plays a crucial role in regulating SL levels. Our study elucidates the structural and regulatory mechanisms of CXE15. We present four crystal structures capturing the conformational dynamics of CXE15, revealing a unique N-terminal extension (Nt) that transitions from a β-sheet in monomers to an intertwined helical structure in dimers. Only the dimeric form is catalytically active, as it forms a hydrophobic cavity for SLs between its two active sites. The moderate dimerisation affinity allows for genetic regulation through protein expression levels. Additionally, we identify an environment-controlled regulation mechanism. Under oxidising conditions, a disulphide bond forms between Cys14 of the two monomers, blocking the active site and inhibiting SL cleavage. This redox-sensitive inhibition of SL catabolism, triggered by reactive oxygen species (ROS) in response to abiotic stress, suggests a mechanism for maintaining high SL levels under beneficial conditions. Our findings provide molecular insights into the regulation of SL homeostasis and catabolism under stress conditions.
- KAUST Center of Excellence for Smart Health, Biological and Environmental Science and Engineering Division, King Abdullah University of Science and Technology (KAUST), Thuwal, Saudi Arabia.
Organizational Affiliation: