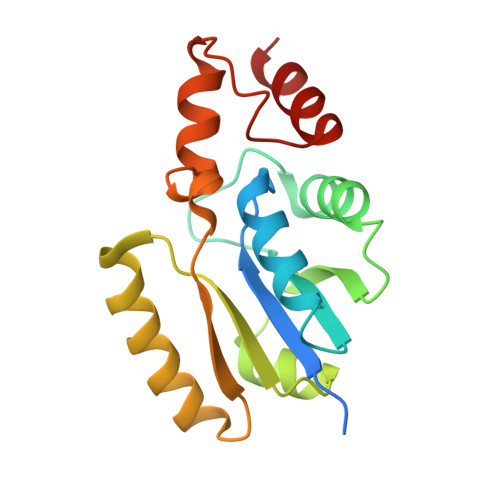
Structure of Phosphopantetheine adenylyltransferase (PPAT) from Enterobacter sp. with the expression tag bound in the substrate binding site of a neighbouring molecule at 2.65 A resolution.
Ahmad, N., Sharma, P., Bhushan, A., Sharma, S., Singh, T.P.To be published.