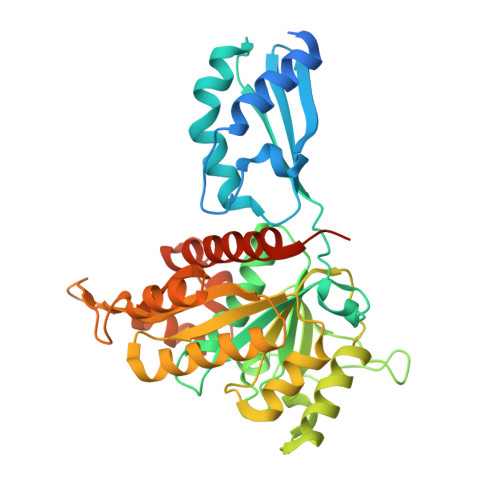
Crystal structure of the iron-sulfur cluster transfer protein ApbC from Escherichia coli.
Yang, J., Duan, Y.F., Liu, L.(2024) Biochem Biophys Res Commun 722: 150167-150167
- PubMed: 38797154 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2024.150167
- Primary Citation Related Structures:
8ZKC - PubMed Abstract:
Iron-sulfur (Fe-S) clusters are ubiquitous and are necessary to sustain basic life processes. The intracellular Fe-S clusters do not form spontaneously and many proteins are required for their biosynthesis and delivery. The bacterial P-loop NTPase family protein ApbC participates in Fe-S cluster assembly and transfers the cluster into apoproteins, with the Walker A motif and CxxC motif being essential for functionality of ApbC in Fe-S protein biogenesis. However, the structural basis underlying the ApbC activity and the motifs' role remains unclear. Here, we report the crystal structure of Escherichia coli ApbC at 2.8 Å resolution. The dimeric structure is in a W shape and the active site is located in the 2-fold center. The function of the motifs can be annotated by structural analyses. ApbC has an additional N-terminal domain that differs from other P-loop NTPases, possibly conferring its inherent specificity in vivo.
- School of Life Sciences, Anhui University, 111 Jiulong Road, Hefei, Anhui, 230601, China. Electronic address: 617113922@qq.com.
Organizational Affiliation: