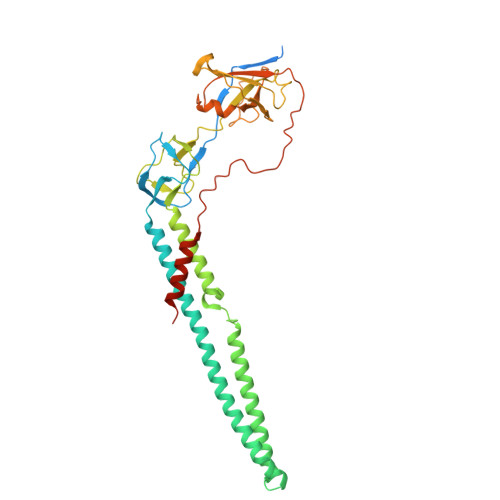
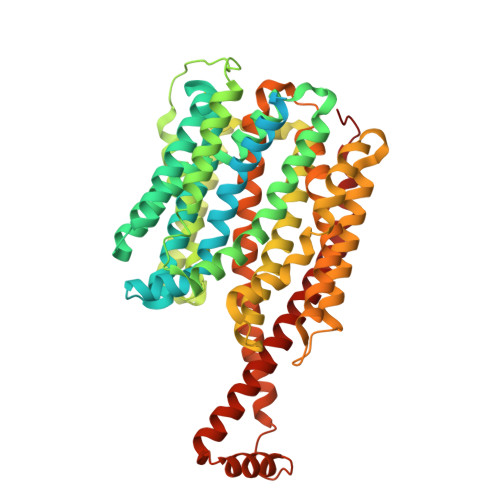
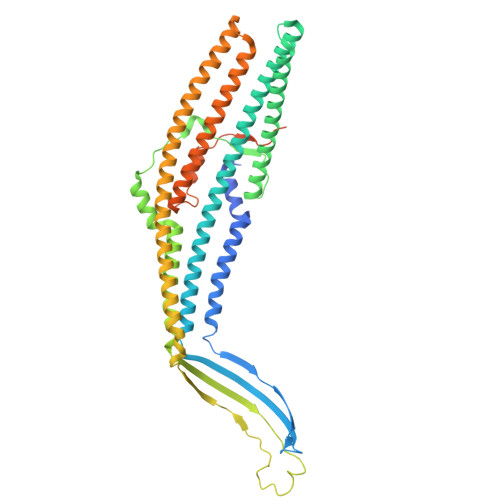
A model for drug transport across two membranes of Gram-negative bacteria by an MFS tripartite assembly.
Zhong, Z., Maimaiti, T., Jackson, M.L., Dong, R., Gao, X., Ouyang, Q., Wang, W., Guo, J., Li, S., Shang, W., Liu, H., Jiang, H., Zhang, S., Zachariae, U., Luisi, B.F., Chao, Y., Du, D.(2026) Nat Commun 17
- PubMed: 41839863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-026-70500-5
- Primary Citation Related Structures:
8ZAL, 8ZAR - PubMed Abstract:
Transport of proteins and small molecules across cellular membrane is crucial for bacterial interaction with the environment and survival against antibiotics. In Gram-negative bacteria that possess two layers of membranes, specialized macromolecular machines are required to transport substrates across the cell envelope, often via an indirect stepwise process. The major facilitator superfamily (MFS)-type tripartite efflux pumps use proton electrochemical gradient to extrude drugs in diverse bacterial species, but the architecture of the assembly and structural mechanisms remain elusive. A representative MFS-type tripartite efflux pump, EmrAB-TolC, mediates resistance to multiple antimicrobial drugs through proton-coupled EmrB, a member of the DHA2 transporter family. Here, we report the high-resolution (3.13 Å) structure of the EmrAB-TolC pump, revealing a distinct, asymmetric architecture emerging from the assembly of TolC:EmrA:EmrB with a ratio of 3:6:1 and contacts that are essential for the pump assembly. Key residues involved in drug transport are identified and corroborated by mutagenesis and antibiotic sensitivity assays. The structural and functional data support a model for one-step drug transport by the MFS pump across the entire envelope of Gram-negative bacteria.
- School of Life Science and Technology, ShanghaiTech University, Shanghai, China.
Organizational Affiliation: