Molecular insights into ligand recognition and signaling of OXGR1.
Liu, A., Liu, Y., Long, Y., Ye, R.D.(2025) Nat Commun 16: 11205-11205
- PubMed: 41413034 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-67101-z
- Primary Citation Related Structures:
8YYW, 8YYX - PubMed Abstract:
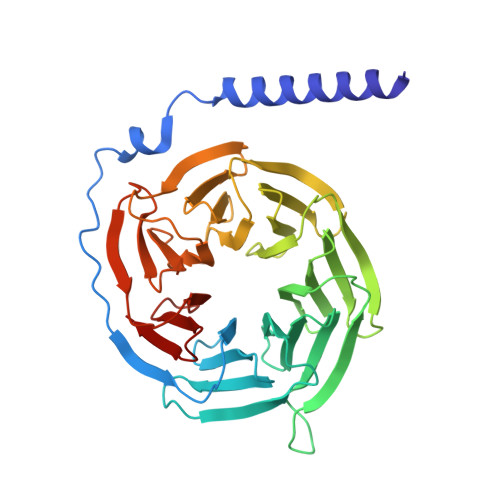
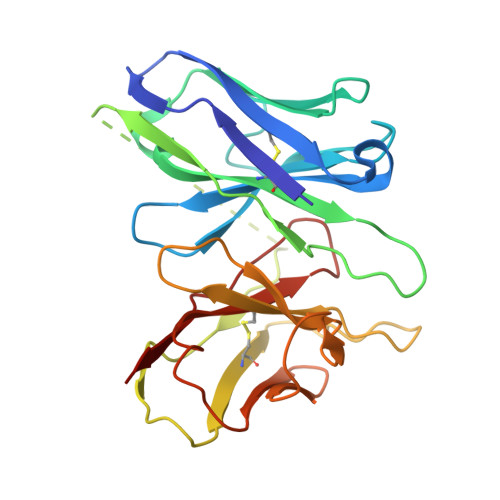
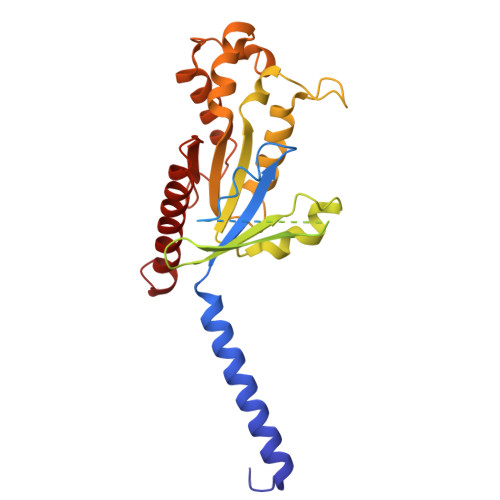
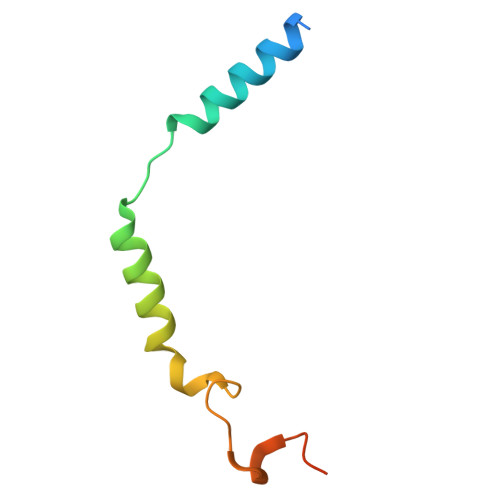
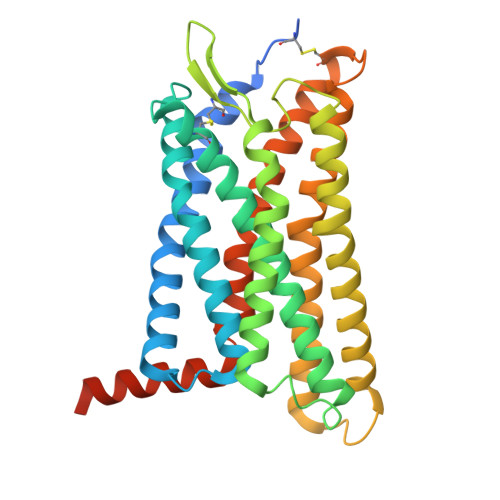
GPR99/OXGR1 is a G protein-coupled receptor (GPCR) with two endogenous agonists, the tricarboxylic acid cycle derivative 2-oxoglutarate (α-ketoglutarate) and the inflammatory mediator cysteinyl leukotriene E4 (LTE 4 ), hence also termed CysLT3 receptor. How GPR99/OXGR1 recognizes two distinct ligands is a biologically important question. Here we present cryo-EM structures of GPR99/OXGR1-Gq complexed with oxoglutarate and LTE 4 , respectively. The oxoglutarate-bound structure shows a binding pocket surrounded by the transmembrane domains (TM), with a primary site and an accessory site for simultaneous binding of two oxoglutarate molecules for full activation of the receptor. The TM binding pocket, however, is too small to accommodate the cysteinyl leukotriene LTE 4 . Alanine substitution of key residues for oxoglutarate binding had little impact on LTE 4 -induced signaling. A distinct site in between TM3/4/5 just above intracellular loop 2 was identified in the solved structure with LTE 4 , but the densities were less well-defined. Alanine substitution of amino acids potentially involved in LTE 4 interaction at this site abrogated LTE 4 -induced receptor activation without affecting oxoglutarate-induced signaling. Both ligands activated GPR99/OXGR1 primarily through the Gq pathway, but LTE 4 also induced Gi signaling. These findings illustrate the structural basis for GPR99/OXGR1 to interact with structurally distict oxoglutarate and LTE 4 .
- The Affiliated Dongguan Songshan Lake Central Hospital, Guangdong Medical University, Dongguan, Guangdong, China. liuaijun@cuhk.edu.cn.
Organizational Affiliation: