KY216-tubulin complex captures VASH2 to inhibit NSCLC metastasis.
He, X.Y., Liu, C.Y., Ding, X.Y., Lin, Q., Leng, J.F., Xiao, C.M., Zang, X., Wang, M.Y., Yin, Y., Kong, L.Y., Xia, Y.Z.(2025) Nat Commun 17: 191-191
- PubMed: 41345086 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66817-2
- Primary Citation Related Structures:
8YRK - PubMed Abstract:
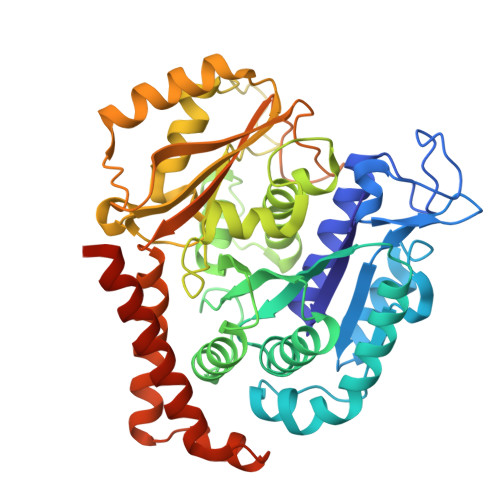
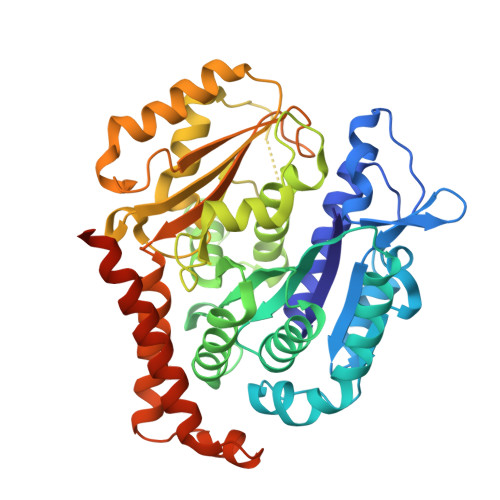
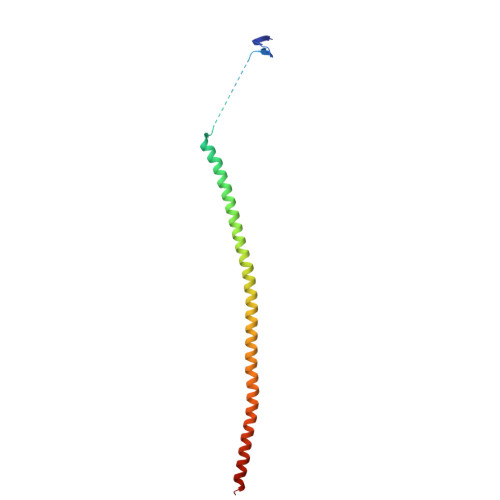
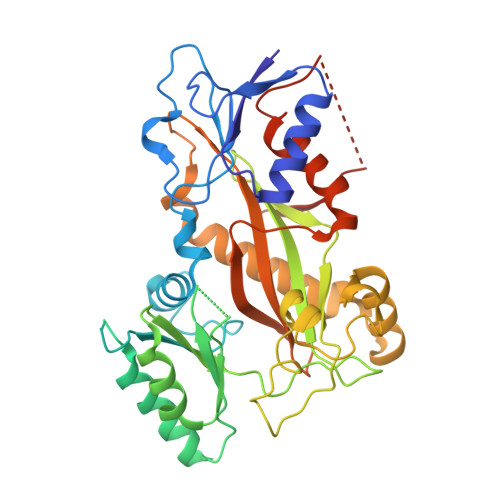
Metastasis significantly impacts the mortality rate of non-small cell lung cancer (NSCLC) patients. Numerous microtubule-targeting agents (MTAs) reveal anti-metastatic efficacy, but the mechanism remains unclear. In this research, we employ KY216, a microtubule inhibitor, to generate a crystal in complex with αβ-tubulin, and illustrate that the KY216-tubulin combination binds vasohibin-2 (VASH2) to restrain NSCLC metastasis. Through crystal structure analysis, specific interaction sites between KY216 and curved tubulin are identified. KY216 decreases VASH2 levels, hindering the epithelial-mesenchymal transition (EMT) process in NSCLC. Moreover, the MTA enhances the binding of VASH2 to α-tubulin, prevents the activation of zinc finger E-box binding homolog 1 (ZEB1) by VASH2, promotes detyrosination of α-tubulin, and ultimately suppresses EMT. Additionally, KY216 elevates the levels of miR-429 to target the 3'-untranslated region (3' UTR) of VASH2 and ZEB1 transcripts and inhibits EMT, at least partially, via the miR-429/VASH2/ZEB1 axis to block NSCLC metastasis. Overall, our investigation offers valuable insights into the roles of MTAs and VASH2 in NSCLC metastasis.
- Basic Medical Research Innovation Center for Anti-Cancer Drugs, MOE and State Key Laboratory of Natural Medicines, China Pharmaceutical University, Nanjing, China.
Organizational Affiliation: