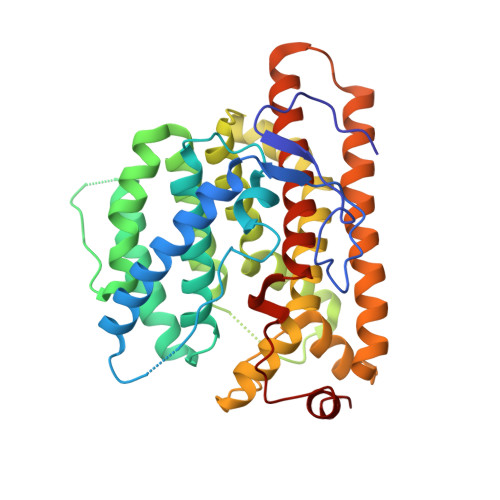
Structural Insights Into the Terpene Cyclization Domains of Two Fungal Sesterterpene Synthases and Enzymatic Engineering for Sesterterpene Diversification.
Xu, M., Xu, H., Lei, Z., Xing, B., Dickschat, J.S., Yang, D., Ma, M.(2024) Angew Chem Int Ed Engl 63: e202405140-e202405140
- PubMed: 38584136 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202405140
- Primary Citation Related Structures:
8YL9, 8YLA - PubMed Abstract:
Little is known about the structures and catalytic mechanisms of sesterterpene synthases (StTSs), which greatly hinders the structure-based engineering of StTSs for structural diversity expansion of sesterterpenes. We here report on the crystal structures of the terpene cyclization (TC) domains of two fungal StTSs: sesterfisherol synthase (NfSS) and sesterbrasiliatriene synthase (PbSS). Both TC structures contain benzyltriethylammonium chloride (BTAC), pyrophosphate (PPi), and magnesium ions (Mg 2+ ), clearly defining the catalytic active sites. A combination of theory and experiments including carbocationic intermediates modeling, site-directed mutagenesis, and isotope labeling provided detailed insights into the structural basis for their catalytic mechanisms. Structure-based engineering of NfSS and PbSS resulted in the formation of 20 sesterterpenes including 13 new compounds and four pairs of epimers with different configurations at C18. These results expand the structural diversity of sesterterpenes and provide important insights for future synthetic biology research.
- State Key Laboratory of Natural and Biomimetic Drugs, School of Pharmaceutical Sciences, Peking University, 38 Xueyuan Road, Haidian District, Beijing, 100191, China.
Organizational Affiliation: