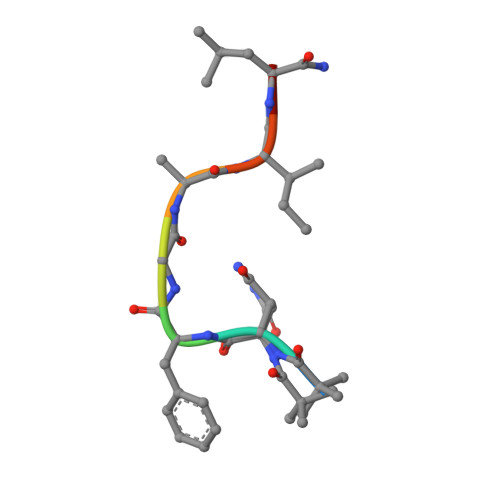
Catalysis driven by an amyloid-substrate complex.
Sawazaki, T., Sasaki, D., Sohma, Y.(2024) Proc Natl Acad Sci U S A 121: e2314704121-e2314704121
- PubMed: 38691589 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2314704121
- Primary Citation Related Structures:
8YG2 - PubMed Abstract:
Amine modification through nucleophilic attack of the amine functionality is a very common chemical transformation. Under biorelevant conditions using acidic-to-neutral pH buffer, however, the nucleophilic reaction of alkyl amines (pKa ≈ 10) is not facile due to the generation of ammonium ions lacking nucleophilicity. Here, we disclose a unique molecular transformation system, c atalysis driven by a myloid- s ubstrate comp l ex (CASL), that promotes amine modifications in acidic buffer. Ammonium ions attached to molecules with amyloid-binding capability were activated through deprotonation due to the close proximity to the amyloid catalyst formed by Ac-Asn-Phe-Gly-Ala-Ile-Leu-NH 2 ( NL6 ), derived from islet amyloid polypeptide (IAPP). Under the CASL conditions, alkyl amines underwent various modifications, i.e., acylation, arylation, cyclization, and alkylation, in acidic buffer. Crystallographic analysis and chemical modification studies of the amyloid catalysts suggested that the carbonyl oxygen of the Phe-Gly amide bond of NL6 plays a key role in activating the substrate amine by forming a hydrogen bond. Using CASL, selective conversion of substrates possessing equivalently reactive amine functionalities was achieved in catalytic reactions using amyloids. CASL provides a unique method for applying nucleophilic conversion reactions of amines in diverse fields of chemistry and biology.
- Department of Medicinal Chemistry, School of Pharmaceutical Sciences, Wakayama Medical University, Wakayama 640-8156, Japan.
Organizational Affiliation: