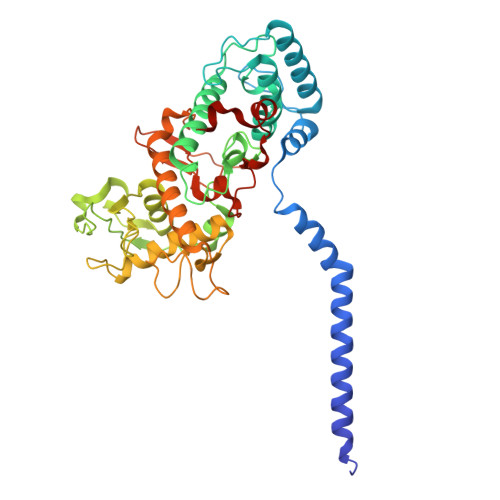
Cryo-EM structure of a minimal reaction center-light-harvesting complex from the phototrophic bacterium Chloroflexus aurantiacus.
Huang, G., Dong, S., Ma, L., Li, L., Ju, J., Wang, M.J., Zhang, J.P., Sui, S.F., Qin, X.(2025) J Integr Plant Biol 67: 967-978
- PubMed: 39912559 Search on PubMed
- DOI: https://doi.org/10.1111/jipb.13853
- Primary Citation Related Structures:
8YDM - PubMed Abstract:
Photosynthetic organisms have developed various light-harvesting antenna systems to capture light and transfer energy to reaction centers (RCs). Simultaneous utilization of the integral membrane light-harvesting antenna (LH complex) and the extrinsic antenna (chlorosomes) makes the phototrophic bacterium Chloroflexus (Cfx.) aurantiacus an ideal model for studying filamentous anoxygenic phototrophs (FAPs). Here, we determined the structure of a minimal RC-LH photocomplex from Cfx. aurantiacus J-10-fl (CaRC-LH) at 3.05-Å resolution. The CaRC-LH binds only to seven LH subunits, which form a crescent-shaped antenna surrounding the movable menaquinone-10 (Q B ) binding site of CaRC. In this complex with minimal LH units, an extra antenna is required to ensure sufficient light-gathering, providing a clear explanation for the presence of chlorosomes in Cfx. aurantiacus. More importantly, the semicircle of the antenna represents a novel RC-LH assembly pattern. Our structure provides a basis for understanding the existence of chlorosomes in Cfx. aurantiacus and the possible assembly pattern of RC-LH.
- State Key Laboratory of Membrane Biology, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing, 100084, China.
Organizational Affiliation: