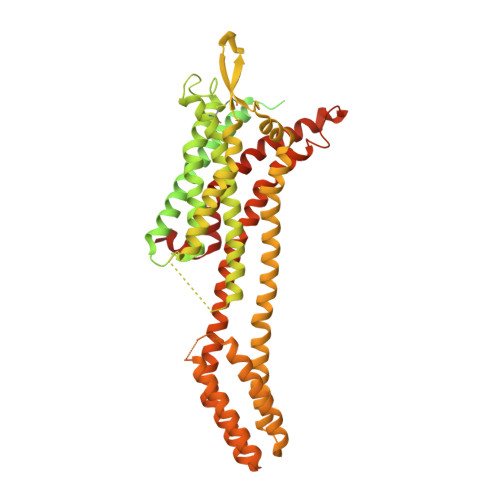Structural basis of antagonist selectivity in endothelin receptors.
Hou, J., Liu, S., Zhang, X., Tu, G., Wu, L., Zhang, Y., Yang, H., Li, X., Liu, J., Jiang, L., Tan, Q., Bai, F., Liu, Z., Miao, C., Hua, T., Luo, Z.(2024) Cell Discov 10: 79-79
- PubMed: 39075075 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-024-00705-9
- Primary Citation Related Structures:
8XVE, 8XVH, 8XVI, 8XVJ, 8XVK, 8XVL - PubMed Abstract:
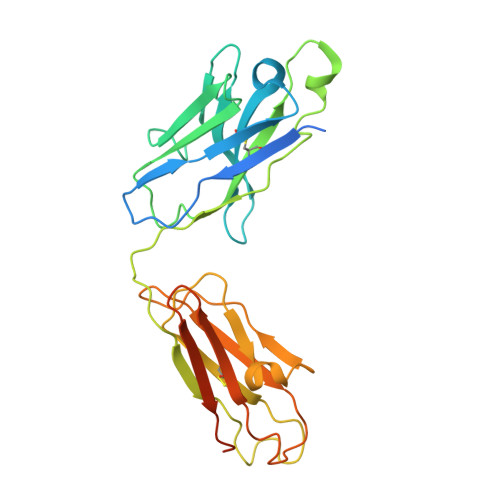
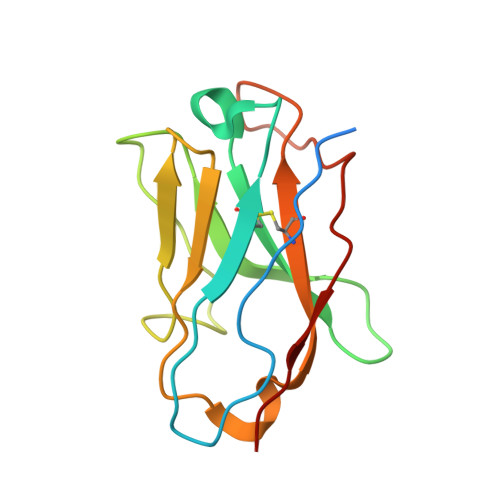
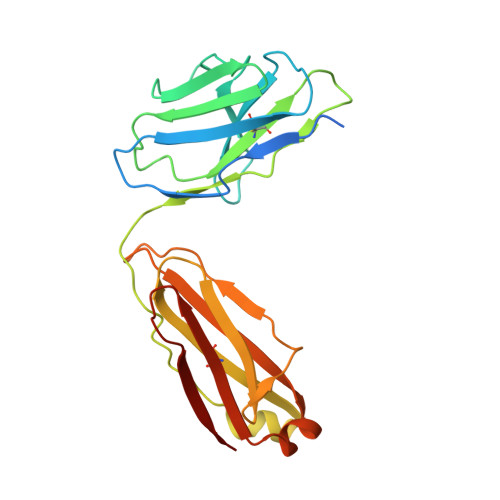
Endothelins and their receptors, ET A and ET B , play vital roles in maintaining vascular homeostasis. Therapeutically targeting endothelin receptors, particularly through ET A antagonists, has shown efficacy in treating pulmonary arterial hypertension (PAH) and other cardiovascular- and renal-related diseases. Here we present cryo-electron microscopy structures of ET A in complex with two PAH drugs, macitentan and ambrisentan, along with zibotentan, a selective ET A antagonist, respectively. Notably, a specialized anti-ET A antibody facilitated the structural elucidation. These structures, together with the active-state structures of ET-1-bound ET A and ET B , and the agonist BQ3020-bound ET B , in complex with G q , unveil the molecular basis of agonist/antagonist binding modes in endothelin receptors. Key residues that confer antagonist selectivity to endothelin receptors were identified along with the activation mechanism of ET A . Furthermore, our results suggest that ECL2 in ET A can serve as an epitope for antibody-mediated receptor antagonism. Collectively, these insights establish a robust theoretical framework for the rational design of small-molecule drugs and antibodies with selective activity against endothelin receptors.
- Cardiac Intensive Care Center, Zhongshan Hospital, Fudan University, Shanghai, China.
Organizational Affiliation: