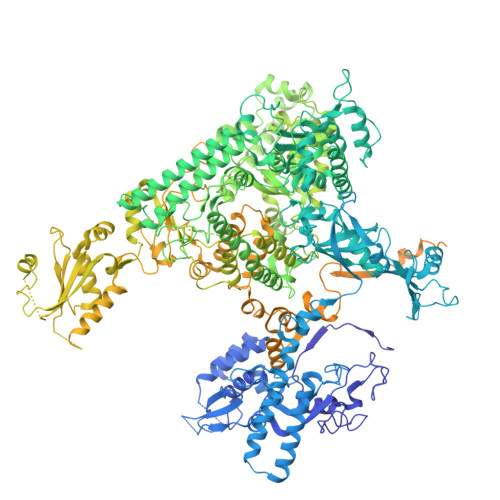
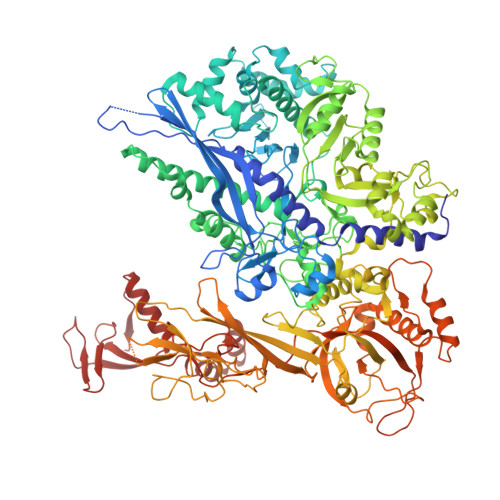
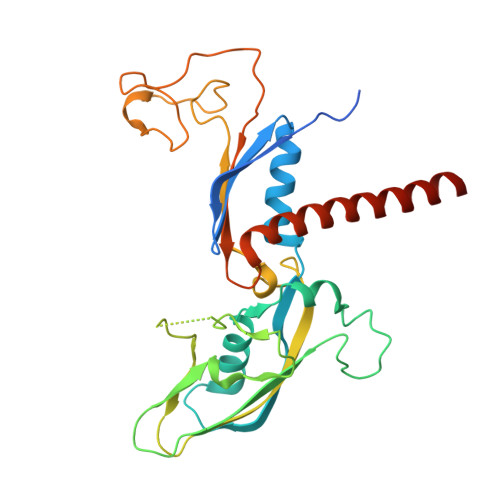
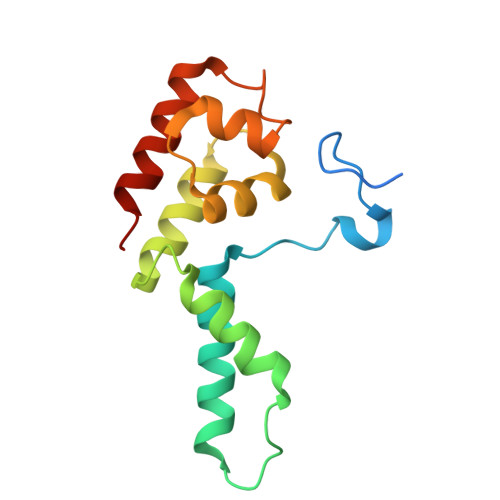
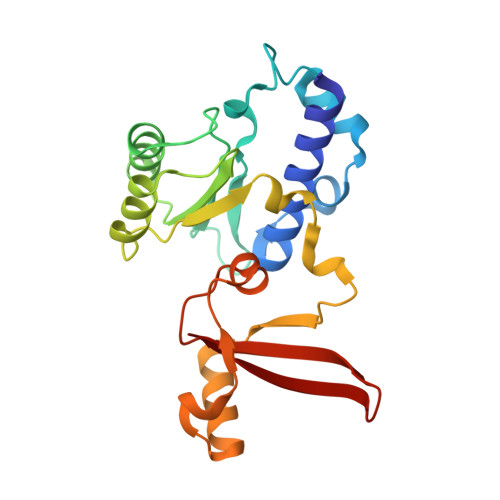
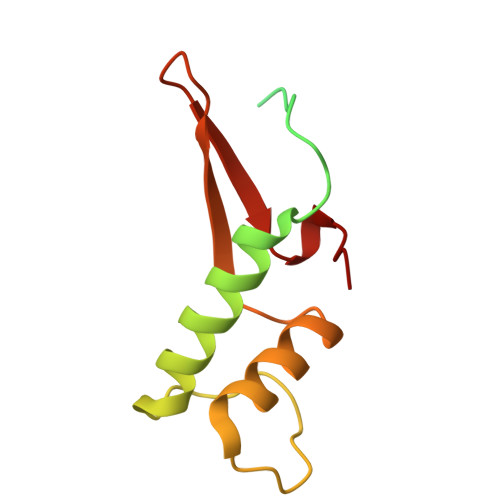
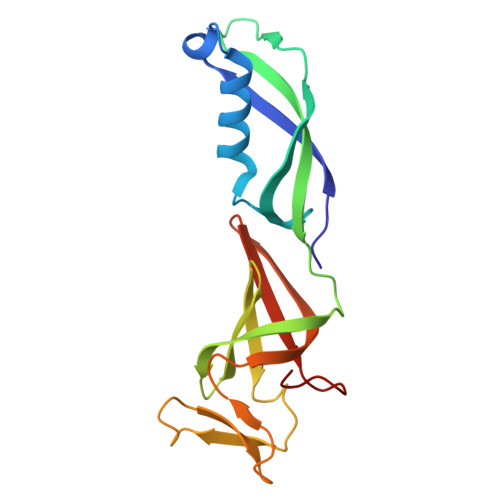
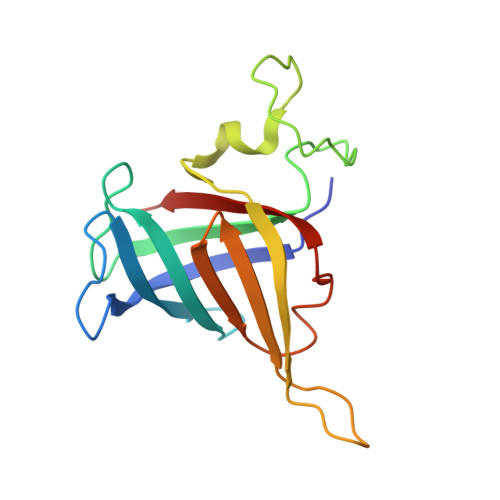
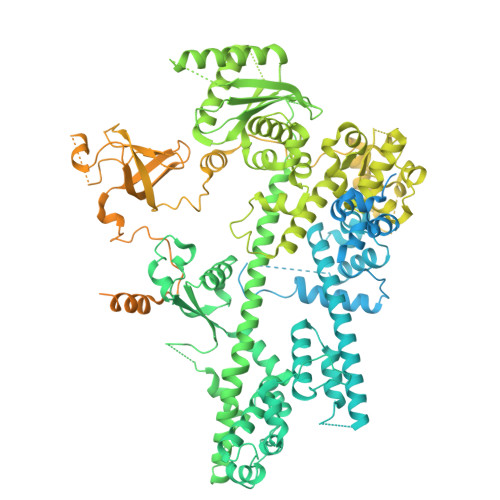
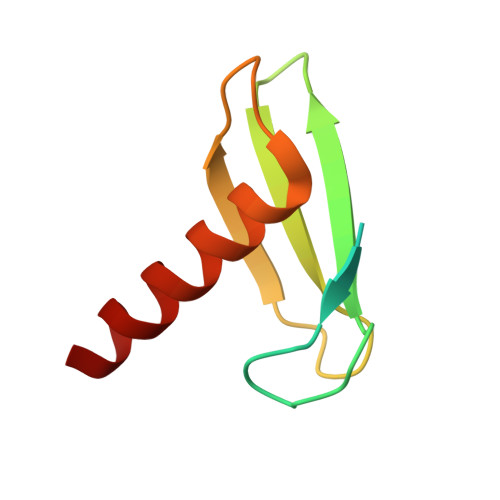
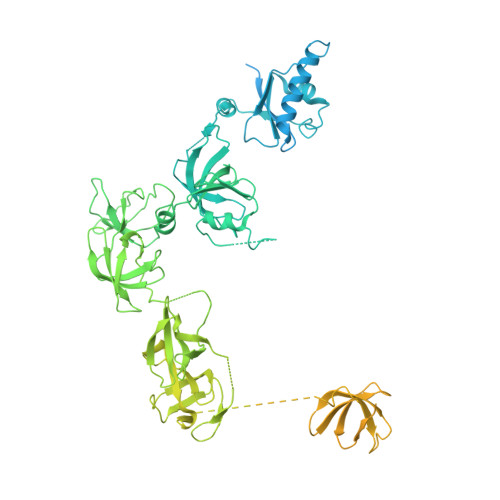
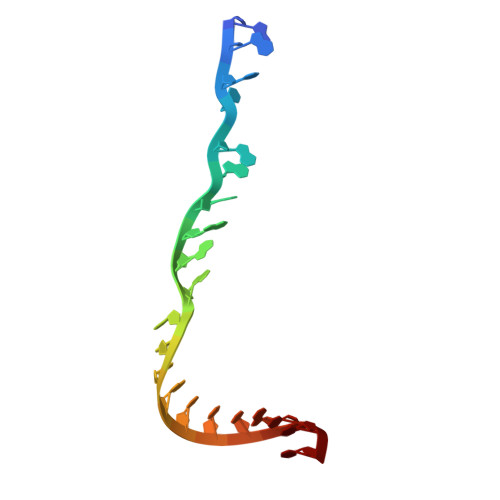
Multiple structures of RNA polymerase II isolated from human nuclei by ChIP-CryoEM analysis.
Kujirai, T., Kato, J., Yamamoto, K., Hirai, S., Fujii, T., Maehara, K., Harada, A., Negishi, L., Ogasawara, M., Yamaguchi, Y., Ohkawa, Y., Takizawa, Y., Kurumizaka, H.(2025) Nat Commun 16: 4724-4724
- PubMed: 40436841 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-59580-x
- Primary Citation Related Structures:
8XRJ, 8XRM, 8XSO, 8XVS - PubMed Abstract:
RNA polymerase II (RNAPII) is a central transcription enzyme that exists as multiple forms with or without accessory factors, and transcribes the genomic DNA packaged in chromatin. To understand how RNAPII functions in the human genome, we isolate transcribing RNAPII complexes from human nuclei by chromatin immunopurification, and determine the cryo-electron microscopy structures of RNAPII elongation complexes (ECs) associated with genomic DNA in distinct forms, without or with the elongation factors SPT4/5, ELOF1, and SPT6. This ChIP-cryoEM method also reveals the two EC-nucleosome complexes corresponding nucleosome disassembly/reassembly processes. In the structure of EC-downstream nucleosome, EC paused at superhelical location (SHL) -5 in the nucleosome, suggesting that SHL(-5) pausing occurs in a sequence-independent manner during nucleosome disassembly. In the structure of the EC-upstream nucleosome, EC directly contacts the nucleosome through the nucleosomal DNA-RPB4/7 stalk and the H2A-H2B dimer-RPB2 wall interactions, suggesting that EC may be paused during nucleosome reassembly. These representative EC structures transcribing the human genome provide mechanistic insights into understanding RNAPII transcription on chromatin.
- Laboratory of Chromatin Structure and Function, Institute for Quantitative Biosciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: