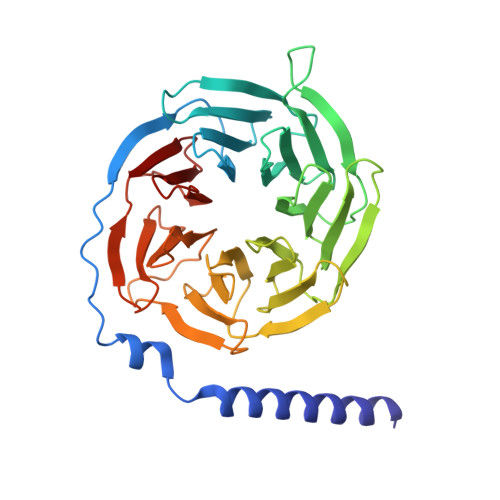
Cryo-EM Structure and Biochemical Analysis of the Human Chemokine Receptor CCR8.
Peng, Q., Jiang, H., Cheng, X., Wang, N., Zhou, S., Zhang, Y., Yang, T., Chen, Y., Zhang, W., Lv, S., Nan, W., Wang, J., Fan, G.H., Li, J., Zhang, J.(2024) Biochemistry 63: 1892-1900
- PubMed: 38985857
- DOI: https://doi.org/10.1021/acs.biochem.4c00121
- Primary Citation of Related Structures:
8XML - PubMed Abstract:
The C-C motif chemokine receptor 8 (CCR8) is a class A G-protein-coupled receptor that has emerged as a promising therapeutic target in cancer and autoimmune diseases. In the present study, we solved the cryo-electron microscopy (cryo-EM) structure of the human CCR8-G i complex in the absence of a ligand at 2.58 Å. Structural analysis and comparison revealed that our apo CCR8 structure undergoes some conformational changes and is similar to that in the CCL1-CCR8 complex structure, indicating an active state. In addition, the key residues of CCR8 involved in the recognition of LMD-009, a potent nonpeptide agonist, were investigated by mutating CCR8 and testing the calcium flux induced by LMD-009-CCR8 interaction. Three mutants of CCR8, Y113 3.32 A, Y172 4.64 A, and E286 7.39 A, showed a dramatically decreased ability in mediating calcium mobilization, indicating their key interaction with LMD-009 and key roles in activation. These structural and biochemical analyses enrich molecular insights into the agonism and activation of CCR8 and will facilitate CCR8-targeted therapy.
- The MOE Basic Research and Innovation Center for the Targeted Therapeutics of Solid Tumors, School of Basic Medical Sciences, Jiangxi Medical College, Nanchang University, Nanchang 330031, China.
Organizational Affiliation: