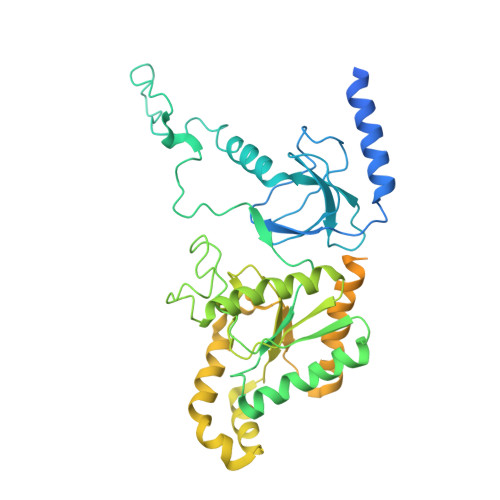
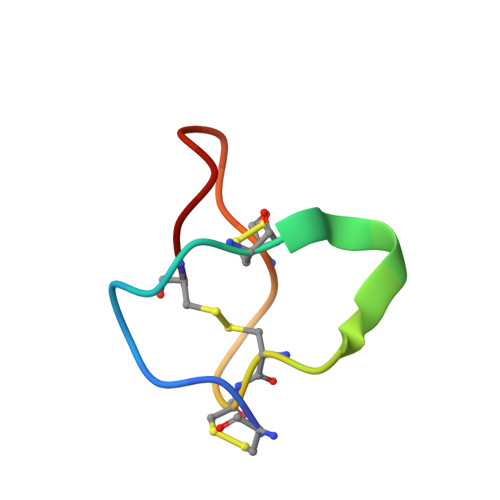
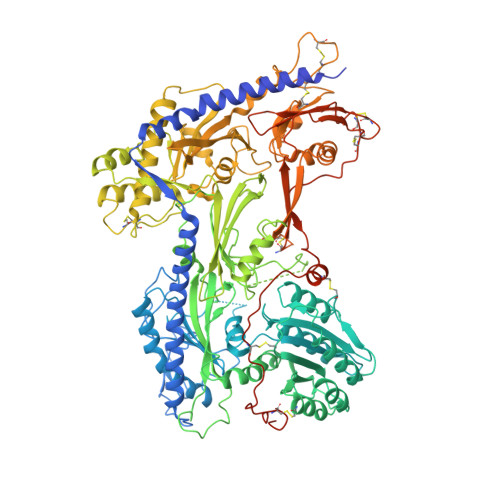
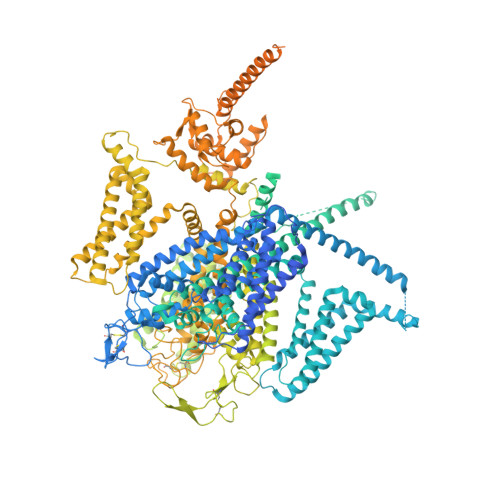
Structural basis for different omega-agatoxin IVA sensitivities of the P-type and Q-type Ca v 2.1 channels.
Li, Z., Cong, Y., Wu, T., Wang, T., Lou, X., Yang, X., Yan, N.(2024) Cell Res 34: 455-457
- PubMed: 38443561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-024-00940-5
- Primary Citation Related Structures:
8X90, 8X91, 8X93 - Beijing Frontier Research Center for Biological Structures, State Key Laboratory of Membrane Biology, Tsinghua-Peking Joint Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing, China. lizhangq@tsinghua.edu.cn.
Organizational Affiliation: