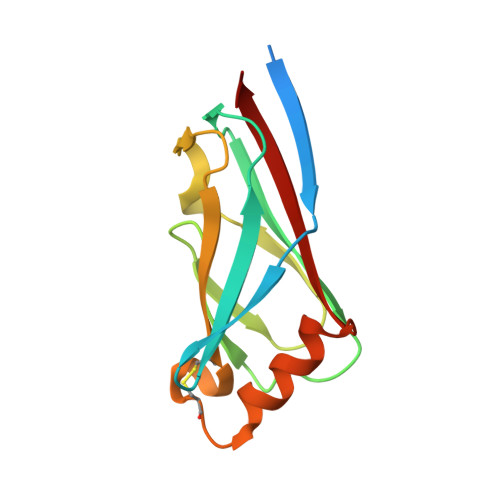
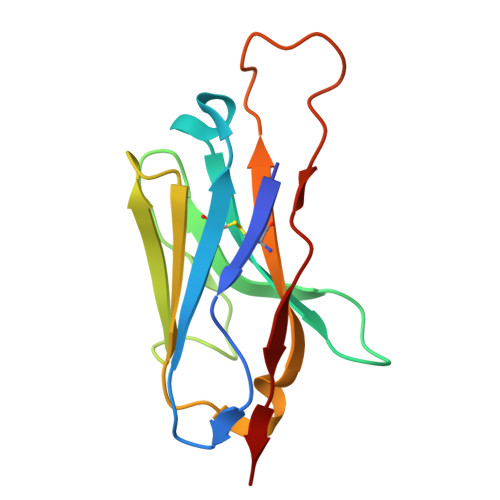
A whole-cell platform for discovering synthetic cell adhesion molecules in bacteria.
Chen, P.Y., Chen, Y.C., Chen, P.P., Lin, K.T., Sargsyan, K., Hsu, C.P., Wang, W.L., Hsia, K.C., Ting, S.Y.(2024) Nat Commun 15: 6568-6568
- PubMed: 39095377 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51017-1
- Primary Citation Related Structures:
8X7N - PubMed Abstract:
Developing programmable bacterial cell-cell adhesion is of significant interest due to its versatile applications. Current methods that rely on presenting cell adhesion molecules (CAMs) on bacterial surfaces are limited by the lack of a generalizable strategy to identify such molecules targeting bacterial membrane proteins in their natural states. Here, we introduce a whole-cell screening platform designed to discover CAMs targeting bacterial membrane proteins within a synthetic bacteria-displayed nanobody library. Leveraging the potency of the bacterial type IV secretion system-a contact-dependent DNA delivery nanomachine-we have established a positive feedback mechanism to selectively enrich for bacteria displaying nanobodies that target antigen-expressing cells. Our platform successfully identified functional CAMs capable of recognizing three distinct outer membrane proteins (TraN, OmpA, OmpC), demonstrating its efficacy in CAM discovery. This approach holds promise for engineering bacterial cell-cell adhesion, such as directing the antibacterial activity of programmed inhibitor cells toward target bacteria in mixed populations.
- Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan.
Organizational Affiliation: