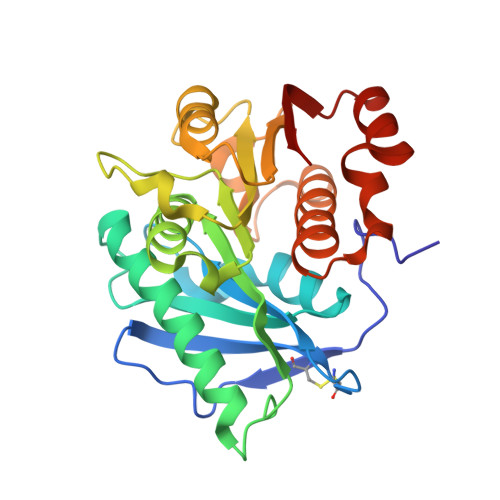
The unique salt bridge network in GlacPETase: a key to its stability.
Qi, X., Wu, Y., Zhang, S.T., Yin, C.F., Ji, M., Liu, Y., Xu, Y., Zhou, N.Y.(2024) Appl Environ Microbiol 90: e0224223-e0224223
- PubMed: 38358247
- DOI: https://doi.org/10.1128/aem.02242-23
- Primary Citation Related Structures:
8X6V - PubMed Abstract:
The extensive accumulation of polyethylene terephthalate (PET) has become a critical environmental issue. PET hydrolases can break down PET into its building blocks. Recently, we identified a glacial PET hydrolase GlacPETase sharing less than 31% amino acid identity with any known PET hydrolases. In this study, the crystal structure of GlacPETase was determined at 1.8 Å resolution, revealing unique structural features including a distinctive N-terminal disulfide bond and a specific salt bridge network. Site-directed mutagenesis demonstrated that the disruption of the N-terminal disulfide bond did not reduce GlacPETase's thermostability or its catalytic activity on PET. However, mutations in the salt bridges resulted in changes in melting temperature ranging from -8°C to +2°C and the activity on PET ranging from 17.5% to 145.5% compared to the wild type. Molecular dynamics simulations revealed that these salt bridges stabilized the GlacPETase's structure by maintaining their surrounding structure. Phylogenetic analysis indicated that GlacPETase represented a distinct branch within PET hydrolases-like proteins, with the salt bridges and disulfide bonds in this branch being relatively conserved. This research contributed to the improvement of our comprehension of the structural mechanisms that dictate the thermostability of PET hydrolases, highlighting the diverse characteristics and adaptability observed within PET hydrolases.IMPORTANCEThe pervasive problem of polyethylene terephthalate (PET) pollution in various terrestrial and marine environments is widely acknowledged and continues to escalate. PET hydrolases, such as GlacPETase in this study, offered a solution for breaking down PET. Its unique origin and less than 31% identity with any known PET hydrolases have driven us to resolve its structure. Here, we report the correlation between its unique structure and biochemical properties, focusing on an N-terminal disulfide bond and specific salt bridges. Through site-directed mutagenesis experiments and molecular dynamics simulations, the roles of the N-terminal disulfide bond and salt bridges were elucidated in GlacPETase. This research enhanced our understanding of the role of salt bridges in the thermostability of PET hydrolases, providing a valuable reference for the future engineering of PET hydrolases.
- State Key Laboratory of Microbial Metabolism, Joint International Research Laboratory of Metabolic and Developmental Sciences, and School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, Shanghai, China.
Organizational Affiliation: