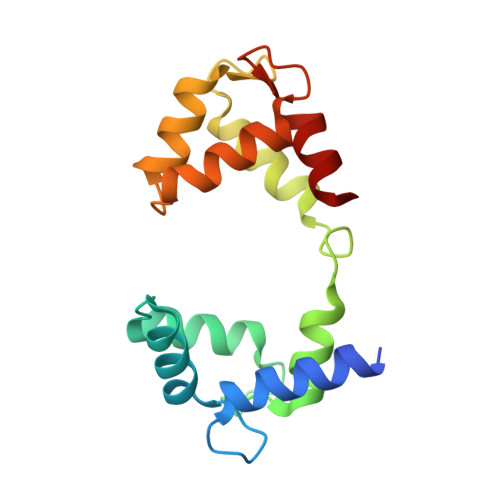
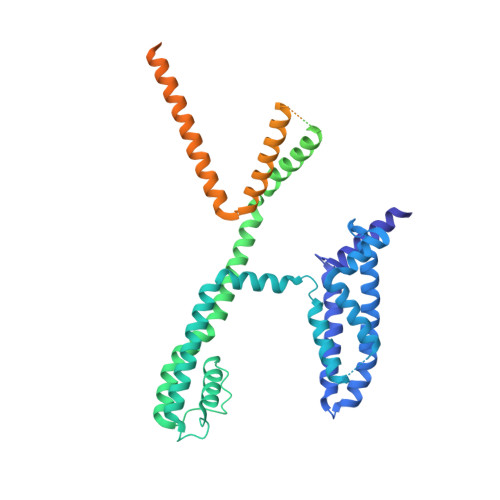A small-molecule activation mechanism that directly opens the KCNQ2 channel.
Zhang, S., Ma, D., Wang, K., Li, Y., Yang, Z., Li, X., Li, J., He, J., Mei, L., Ye, Y., Chen, Z., Shen, J., Hou, P., Guo, J., Zhang, Q., Yang, H.(2024) Nat Chem Biol 20: 847-856
- PubMed: 38167918 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01515-y
- Primary Citation Related Structures:
8IJK, 8X43 - PubMed Abstract:
Pharmacological activation of voltage-gated ion channels by ligands serves as the basis for therapy and mainly involves a classic gating mechanism that augments the native voltage-dependent open probability. Through structure-based virtual screening, we identified a new scaffold compound, Ebio1, serving as a potent and subtype-selective activator for the voltage-gated potassium channel KCNQ2 and featuring a new activation mechanism. Single-channel patch-clamp, cryogenic-electron microscopy and molecular dynamic simulations, along with chemical derivatives, reveal that Ebio1 engages the KCNQ2 activation by generating an extended channel gate with a larger conductance at the saturating voltage (+50 mV). This mechanism is different from the previously observed activation mechanism of ligands on voltage-gated ion channels. Ebio1 caused S6 helices from residues S303 and F305 to perform a twist-to-open movement, which was sufficient to open the KCNQ2 gate. Overall, our findings provide mechanistic insights into the activation of KCNQ2 channel by Ebio1 and lend support for KCNQ-related drug development.
- Shanghai Key Laboratory of Regulatory Biology, Institute of Biomedical Sciences and School of Life Sciences, East China Normal University, Shanghai, China.
Organizational Affiliation: