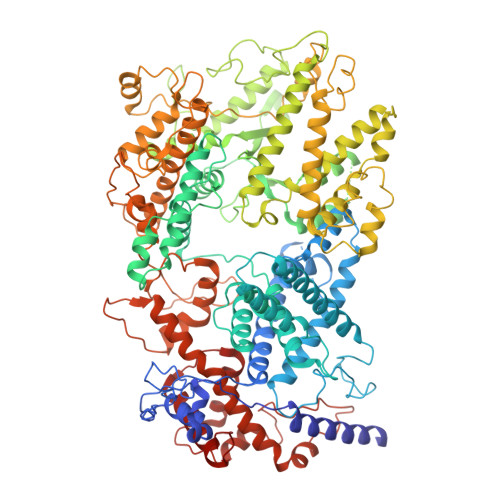
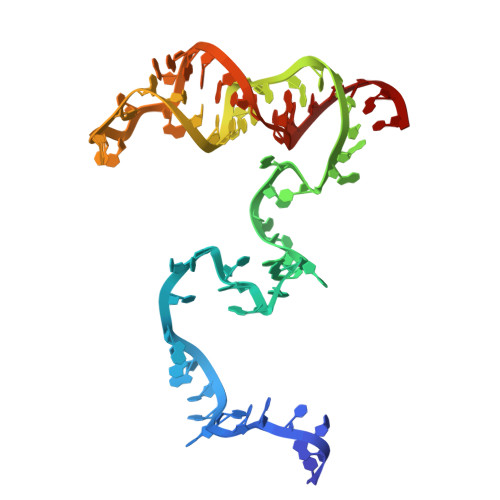
Molecular mechanism for target RNA recognition and cleavage of Cas13h.
Chen, F., Zhang, C., Xue, J., Wang, F., Li, Z.(2024) Nucleic Acids Res 52: 7279-7291
- PubMed: 38661236 Search on PubMed
- DOI: https://doi.org/10.1093/nar/gkae324
- Primary Citation Related Structures:
8WCE, 8WCS - PubMed Abstract:
RNA-targeting type VI CRISPR-Cas effectors are widely used in RNA applications. Cas13h is a recently identified subtype of Cas13 ribonuclease, with strong RNA cleavage activity and robust in vivo RNA knockdown efficiency. However, little is known regarding its biochemical properties and working mechanisms. Biochemical characterization of Cas13h1 indicated that it lacks in vitro pre-crRNA processing activity and adopts a central seed. The cleavage activity of Cas13h1 is enhanced by a R(G/A) 5'-PFS, and inhibited by tag:anti-tag RNA pairing. We determined the structures of Cas13h1-crRNA binary complex at 3.1 Å and Cas13h1-crRNA-target RNA ternary complex at 3.0 Å. The ternary complex adopts an elongated architecture, and encodes a nucleotide-binding pocket within Helical-2 domain to recognize the guanosine at the 5'-end of the target RNA. Base pairing between crRNA guide and target RNA disrupts Cas13h1-guide interactions, leading to dramatic movement of HEPN domains. Upon target RNA engagement, Cas13h1 adopts a complicated activation mechanism, including separation of HEPN catalytic residues and destabilization of the active site loop and NTD domain, to get activated. Collectively, these insights expand our understanding into Cas13 effectors.
- State Key Laboratory of Biocatalysis and Enzyme Engineering, School of Life Sciences, Hubei University, Wuhan, Hubei 430062, China.
Organizational Affiliation: