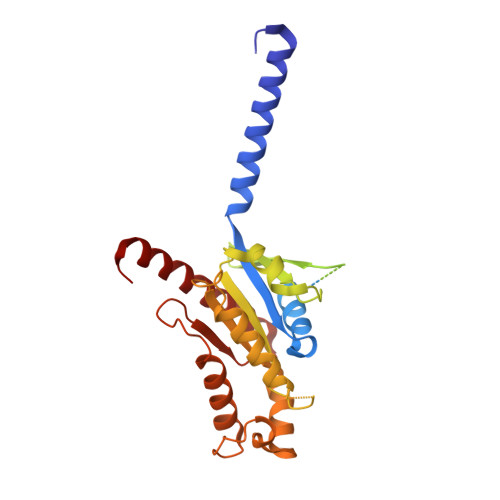
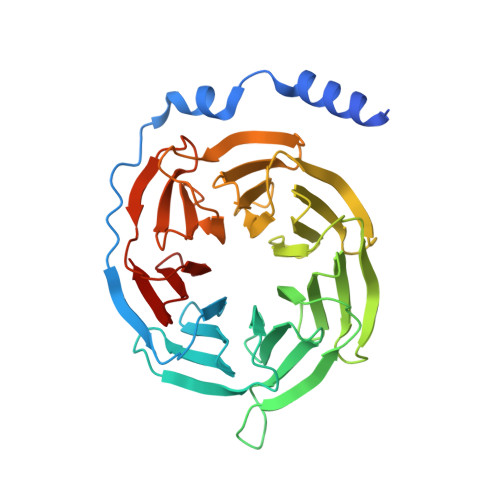
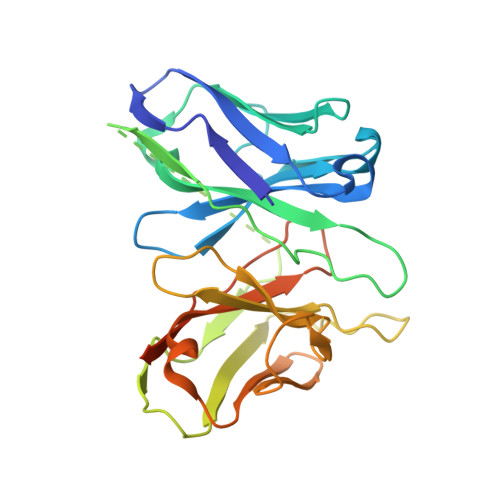
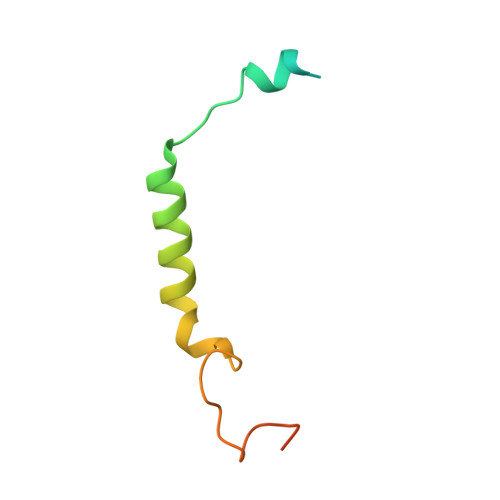
Recognition of methamphetamine and other amines by trace amine receptor TAAR1.
Liu, H., Zheng, Y., Wang, Y., Wang, Y., He, X., Xu, P., Huang, S., Yuan, Q., Zhang, X., Wang, L., Jiang, K., Chen, H., Li, Z., Liu, W., Wang, S., Xu, H.E., Xu, F.(2023) Nature 624: 663-671
- PubMed: 37935377 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06775-1
- Primary Citation Related Structures:
8W87, 8W88, 8W89, 8W8A, 8W8B - PubMed Abstract:
Trace amine-associated receptor 1 (TAAR1), the founding member of a nine-member family of trace amine receptors, is responsible for recognizing a range of biogenic amines in the brain, including the endogenous β-phenylethylamine (β-PEA) 1 as well as methamphetamine 2 , an abused substance that has posed a severe threat to human health and society 3 . Given its unique physiological role in the brain, TAAR1 is also an emerging target for a range of neurological disorders including schizophrenia, depression and drug addiction 2,4,5 . Here we report structures of human TAAR1-G-protein complexes bound to methamphetamine and β-PEA as well as complexes bound to RO5256390, a TAAR1-selective agonist, and SEP-363856, a clinical-stage dual agonist for TAAR1 and serotonin receptor 5-HT 1A R (refs. 6,7 ). Together with systematic mutagenesis and functional studies, the structures reveal the molecular basis of methamphetamine recognition and underlying mechanisms of ligand selectivity and polypharmacology between TAAR1 and other monoamine receptors. We identify a lid-like extracellular loop 2 helix/loop structure and a hydrogen-bonding network in the ligand-binding pockets, which may contribute to the ligand recognition in TAAR1. These findings shed light on the ligand recognition mode and activation mechanism for TAAR1 and should guide the development of next-generation therapeutics for drug addiction and various neurological disorders.
- The State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: