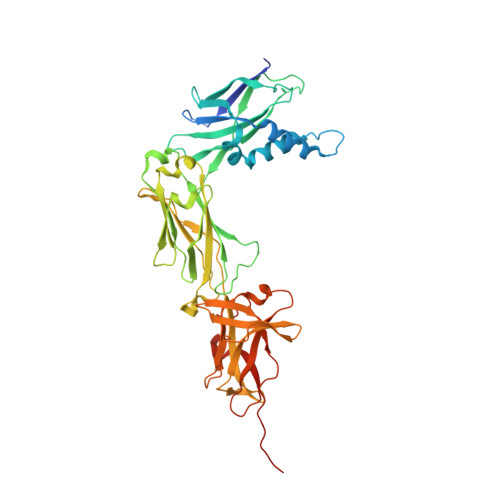
The crystal structure of the N-terminal domain of the backbone pilin LrpA reveals a new closure-and-twist motion for assembling dynamic pili in Ligilactobacillus ruminis.
Prajapati, A., Palva, A., von Ossowski, I., Krishnan, V.(2024) Acta Crystallogr D Struct Biol 80: 474-492
- PubMed: 38935340 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798324005114
- Primary Citation Related Structures:
8KB2, 8KCL, 8KG4, 8W5B, 8WB8 - PubMed Abstract:
Sortase-dependent pili are long surface appendages that mediate attachment, colonization and biofilm formation in certain genera and species of Gram-positive bacteria. Ligilactobacillus ruminis is an autochthonous gut commensal that relies on sortase-dependent LrpCBA pili for host adherence and persistence. X-ray crystal structure snapshots of the backbone pilin LrpA were captured in two atypical bent conformations leading to a zigzag morphology in the LrpCBA pilus structure. Small-angle X-ray scattering and structural analysis revealed that LrpA also adopts the typical linear conformation, resulting in an elongated pilus morphology. Various conformational analyses and biophysical experiments helped to demonstrate that a hinge region located at the end of the flexible N-terminal domain of LrpA facilitates a new closure-and-twist motion for assembling dynamic pili during the assembly process and host attachment. Further, the incongruent combination of flexible domain-driven conformational dynamics and rigid isopeptide bond-driven stability observed in the LrpCBA pilus might also extend to the sortase-dependent pili of other bacteria colonizing a host.
- Laboratory of Structural Microbiology, Regional Centre for Biotechnology, NCR, Biotech Science Cluster, Faridabad 121 001, India.
Organizational Affiliation: