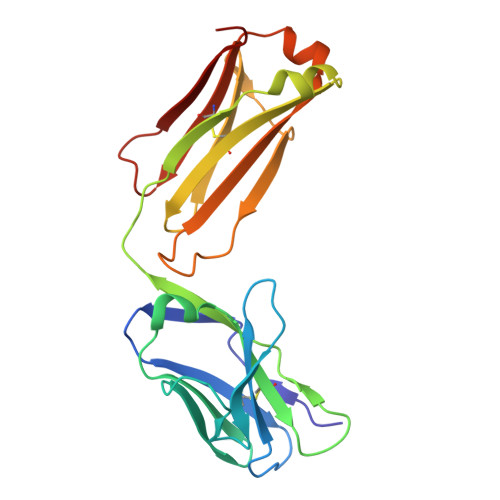
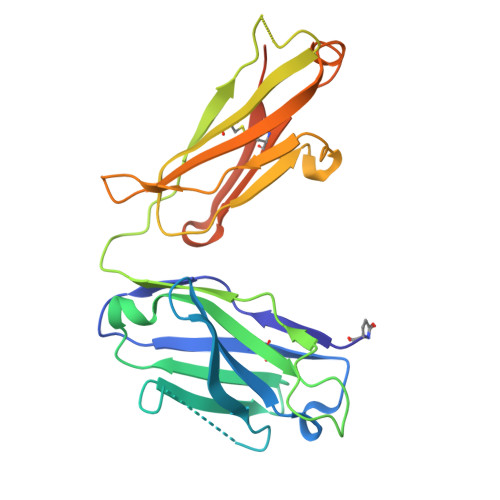
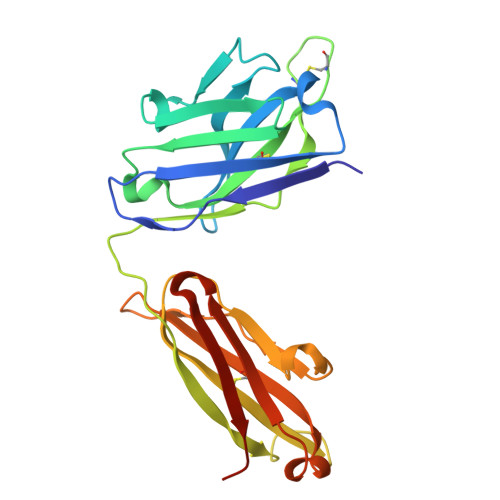
Targeting RSV-neutralizing B cell receptors with anti-idiotypic antibodies.
Scharffenberger, S.C., Wan, Y.H., Homad, L.J., Kher, G., Haynes, A.M., Poudel, B., Sinha, I.R., Aldridge, N., Pai, A., Bibby, M., Chhan, C.B., Davis, A.R., Moodie, Z., Palacio, M.B., Escolano, A., McElrath, M.J., Boonyaratanakornkit, J., Pancera, M., McGuire, A.T.(2024) Cell Rep 43: 114811-114811
- PubMed: 39383036
- DOI: https://doi.org/10.1016/j.celrep.2024.114811
- Primary Citation Related Structures:
8VS7, 8VS8 - PubMed Abstract:
Respiratory syncytial virus (RSV) causes lower respiratory tract infections with significant morbidity and mortality at the extremes of age. Vaccines based on the viral fusion protein are approved for adults over 60, but infant protection relies on passive immunity via antibody transfer or maternal vaccination. An infant vaccine that rapidly elicits protective antibodies would fulfill a critical unmet need. Antibodies arising from the VH3-21/VL1-40 gene pairing can neutralize RSV without the need for affinity maturation, making them attractive to target through vaccination. Here, we develop an anti-idiotypic monoclonal antibody (ai-mAb) immunogen that is specific for unmutated VH3-21/VL1-40 B cell receptors (BCRs). The ai-mAb efficiently engages B cells with bona fide target BCRs and does not activate off-target non-neutralizing B cells, unlike recombinant pre-fusion (preF) protein used in current RSV vaccines. These results establish proof of concept for using an ai-mAb-derived vaccine to target B cells hardwired to produce RSV-neutralizing antibodies.
- Vaccine and Infectious Disease Division, Fred Hutchinson Cancer Center, Seattle, WA 98109, USA; Department of Laboratory Medicine and Pathology, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: