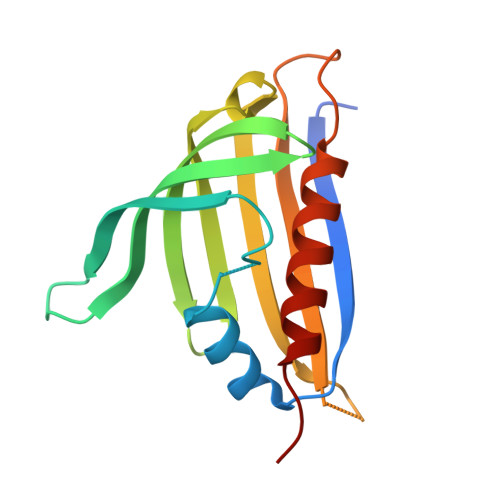
Structural analysis of a ligand-triggered intermolecular disulfide switch in a major latex protein from opium poppy.
Carr, S.C., Facchini, P.J., Ng, K.K.S.(2024) Acta Crystallogr D Struct Biol 80: 675-685
- PubMed: 39207895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798324007733
- Primary Citation Related Structures:
8VO1, 8VO2, 8VO3 - PubMed Abstract:
Several proteins from plant pathogenesis-related family 10 (PR10) are highly abundant in the latex of opium poppy and have recently been shown to play diverse and important roles in the biosynthesis of benzylisoquinoline alkaloids (BIAs). The recent determination of the first crystal structures of PR10-10 showed how large conformational changes in a surface loop and adjacent β-strand are coupled to the binding of BIA compounds to the central hydrophobic binding pocket. A more detailed analysis of these conformational changes is now reported to further clarify how ligand binding is coupled to the formation and cleavage of an intermolecular disulfide bond that is only sterically allowed when the BIA binding pocket is empty. To decouple ligand binding from disulfide-bond formation, each of the two highly conserved cysteine residues (Cys59 and Cys155) in PR10-10 was replaced with serine using site-directed mutagenesis. Crystal structures of the Cys59Ser mutant were determined in the presence of papaverine and in the absence of exogenous BIA compounds. A crystal structure of the Cys155Ser mutant was also determined in the absence of exogenous BIA compounds. All three of these crystal structures reveal conformations similar to that of wild-type PR10-10 with bound BIA compounds. In the absence of exogenous BIA compounds, the Cys59Ser and Cys155Ser mutants appear to bind an unidentified ligand or mixture of ligands that was presumably introduced during expression of the proteins in Escherichia coli. The analysis of conformational changes triggered by the binding of BIA compounds suggests a molecular mechanism coupling ligand binding to the disruption of an intermolecular disulfide bond. This mechanism may be involved in the regulation of biosynthetic reactions in plants and possibly other organisms.
- Department of Biological Sciences, University of Calgary, Calgary, Alberta T2N 1N4, Canada.
Organizational Affiliation: