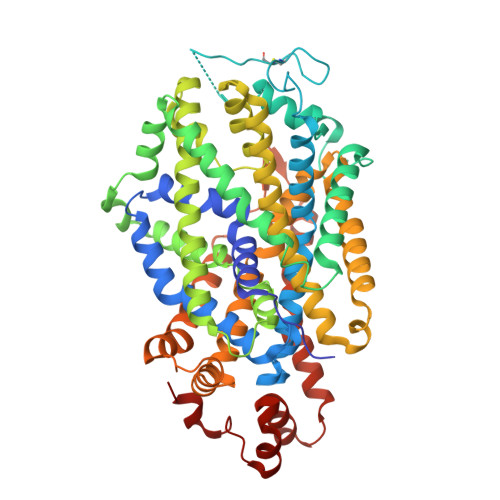
Structure of the human dopamine transporter and mechanisms of inhibition.
Srivastava, D.K., Navratna, V., Tosh, D.K., Chinn, A., Sk, M.F., Tajkhorshid, E., Jacobson, K.A., Gouaux, E.(2024) Nature 632: 672-677
- PubMed: 39112705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07739-9
- Primary Citation Related Structures:
8VBY - PubMed Abstract:
The neurotransmitter dopamine has central roles in mood, appetite, arousal and movement 1 . Despite its importance in brain physiology and function, and as a target for illicit and therapeutic drugs, the human dopamine transporter (hDAT) and mechanisms by which it is inhibited by small molecules and Zn 2+ are without a high-resolution structural context. Here we determine the structure of hDAT in a tripartite complex with the competitive inhibitor and cocaine analogue, (-)-2-β-carbomethoxy-3-β-(4-fluorophenyl)tropane 2 (β-CFT), the non-competitive inhibitor MRS7292 3 and Zn 2 + (ref. 4 ). We show how β-CFT occupies the central site, approximately halfway across the membrane, stabilizing the transporter in an outward-open conformation. MRS7292 binds to a structurally uncharacterized allosteric site, adjacent to the extracellular vestibule, sequestered underneath the extracellular loop 4 (EL4) and adjacent to transmembrane helix 1b (TM1b), acting as a wedge, precluding movement of TM1b and closure of the extracellular gate. A Zn 2+ ion further stabilizes the outward-facing conformation by coupling EL4 to EL2, TM7 and TM8, thus providing specific insights into how Zn 2+ restrains the movement of EL4 relative to EL2 and inhibits transport activity.
- Vollum Institute, Oregon Health and Science University, Portland, OR, USA.
Organizational Affiliation: