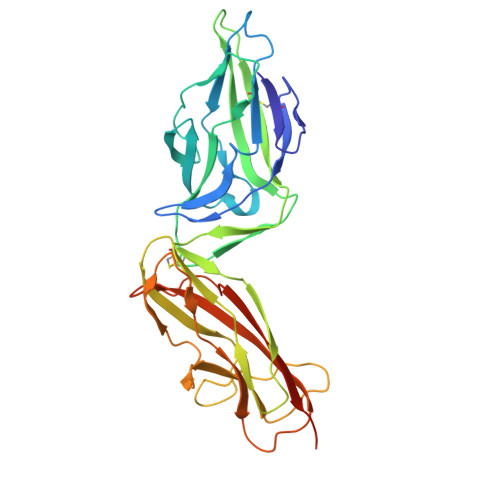
Structure-based design of an immunogenic, conformationally stabilized FimH antigen for a urinary tract infection vaccine.
Silmon de Monerri, N.C., Che, Y., Lees, J.A., Jasti, J., Wu, H., Griffor, M.C., Kodali, S., Hawkins, J.C., Lypowy, J., Ponce, C., Curley, K., Esadze, A., Carcamo, J., McLellan, T., Keeney, D., Illenberger, A., Matsuka, Y.V., Shanker, S., Chorro, L., Gribenko, A.V., Han, S., Anderson, A.S., Donald, R.G.K.(2025) PLoS Pathog 21: e1012325-e1012325
- PubMed: 39970181 Search on PubMed
- DOI: https://doi.org/10.1371/journal.ppat.1012325
- Primary Citation Related Structures:
8V3J, 8V93, 9D6F - PubMed Abstract:
Adhesion of E. coli to the urinary tract epithelium is a critical step in establishing urinary tract infections. FimH is an adhesin positioned on the fimbrial tip which binds to mannosylated proteins on the urinary tract epithelium via its lectin domain (FimHLD). FimH is of interest as a target of vaccines to prevent urinary tract infections (UTI). Previously, difficulties in obtaining purified recombinant FimH from E. coli along with the poor inherent immunogenicity of FimH have hindered the development of effective FimH vaccine candidates. To overcome these challenges, we have devised a novel production method using mammalian cells to produce high yields of homogeneous FimH protein with comparable biochemical and immunogenic properties to FimH produced in E. coli. Next, to optimize conformational stability and immunogenicity of FimH, we used a computational approach to design improved FimH mutants and evaluated their biophysical and biochemical properties, and murine immunogenicity using a bacterial adhesion inhibition assay. This approach identified an immunogenic FimH variant (FimH-donor-strand complemented with FimG peptide 'triple mutant', FimH-DSG TM) capable of blocking bacterial adhesion that is produced at high yields in mammalian cells. By x-ray crystallography, we confirmed that the stabilized structure of the FimHLD in FimH-DSG TM is similar to native FimH on the fimbrial tip. Characterization of monoclonal antibodies elicited by FimH-DSG TM that can block bacterial binding to mannosylated surfaces identified 4 non-overlapping binding sites whose epitopes were mapped via a combinatorial cryogenic electron microscopy approach. Novel inhibitory epitopes in the lectin binding FimH were identified, revealing diverse functional mechanisms of FimH-directed antibodies with relevance to FimH-targeted UTI vaccines.
- Vaccine Research and Development, Pfizer Inc, Pearl River, New York, New York, United States of America.
Organizational Affiliation: