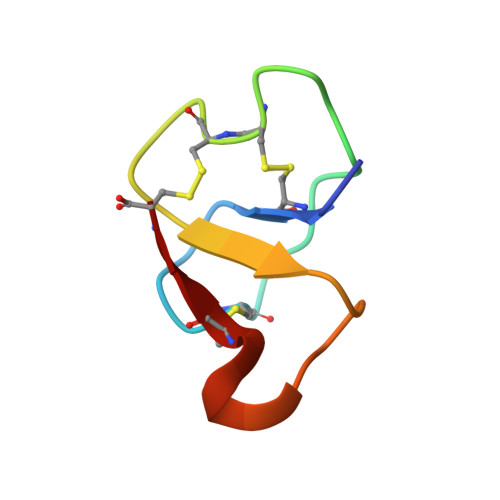
Picking the tyrosine-lock: chemical synthesis of the tyrosyl-DNA phosphodiesterase I inhibitor recifin A and analogues.
Smallwood, T.B., Krumpe, L.R.H., Payne, C.D., Klein, V.G., O'Keefe, B.R., Clark, R.J., Schroeder, C.I., Rosengren, K.J.(2024) Chem Sci 15: 13227-13233
- PubMed: 39183914 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d4sc01976h
- Primary Citation Related Structures:
8V2V - PubMed Abstract:
The peptide recifin A is the inaugural member of the structurally intriguing new fold referred to as a tyrosine-lock. Its central four stranded β-sheet is stabilized by a unique arrangement in which three disulfide bonds and their interconnecting backbone form a ring that wraps around one of the strands, resulting in a Tyr side chain being buried in the molecular core. Here we aimed to establish a synthetic route to this complex class of natural products. Full length recifin A was successfully generated through native chemical ligation chemistry joining two 21 amino acid residue fragments. Surprisingly, reduced linear recifin A readily adopts the correct, topologically-complex fold via random oxidation of the cysteines, suggesting it is highly energetically favored. Utilizing our synthetic strategy, we generated five recifin A analogues to investigate the structural role of the central Tyr residue and provide the first insights into the structure activity relationship of recifin A towards its cancer target tyrosyl-DNA phosphodiesterase I.
- The University of Queensland, School of Biomedical Sciences Brisbane QLD 4072 Australia richard.clark@uq.edu.au j.rosengren@uq.edu.au.
Organizational Affiliation: