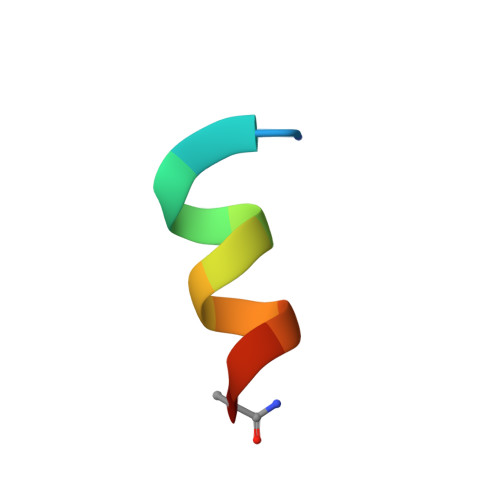Bioinformatics leading to conveniently accessible, helix enforcing, bicyclic ASX motif mimics (BAMMs).
Mi, T., Nguyen, D., Gao, Z., Burgess, K.(2024) Nat Commun 15: 4217-4217
- PubMed: 38760359 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48323-z
- Primary Citation Related Structures:
8UN8, 8UTX - PubMed Abstract:
Helix mimicry provides probes to perturb protein-protein interactions (PPIs). Helical conformations can be stabilized by joining side chains of non-terminal residues (stapling) or via capping fragments. Nature exclusively uses capping, but synthetic helical mimics are heavily biased towards stapling. This study comprises: (i) creation of a searchable database of unique helical N-caps (ASX motifs, a protein structural motif with two intramolecular hydrogen-bonds between aspartic acid/asparagine and following residues); (ii) testing trends observed in this database using linear peptides comprising only canonical L-amino acids; and, (iii) novel synthetic N-caps for helical interface mimicry. Here we show many natural ASX motifs comprise hydrophobic triangles, validate their effect in linear peptides, and further develop a biomimetic of them, Bicyclic ASX Motif Mimics (BAMMs). BAMMs are powerful helix inducing motifs. They are synthetically accessible, and potentially useful to a broad section of the community studying disruption of PPIs using secondary structure mimics.
- Department of Chemistry, Texas A & M University, College Station, TX, 77842, USA.
Organizational Affiliation: