An alphacoronavirus polymerase structure reveals conserved co-factor functions.
Anderson, T.K., Hoferle, P.J., Lee, K.W., Coon, J.J., Kirchdoerfer, R.N.(2023) bioRxiv
- PubMed: 36993498 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2023.03.15.532841
- Primary Citation Related Structures:
8G6R, 8URB - PubMed Abstract:
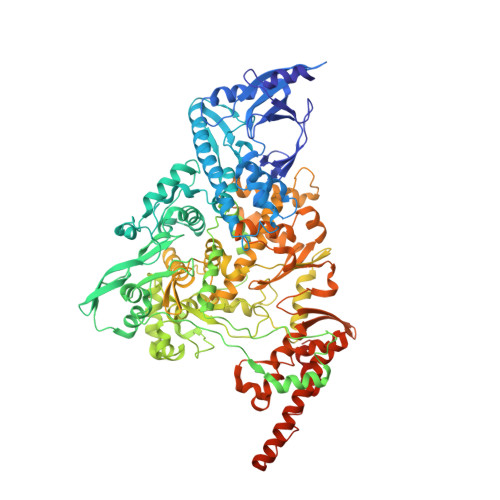
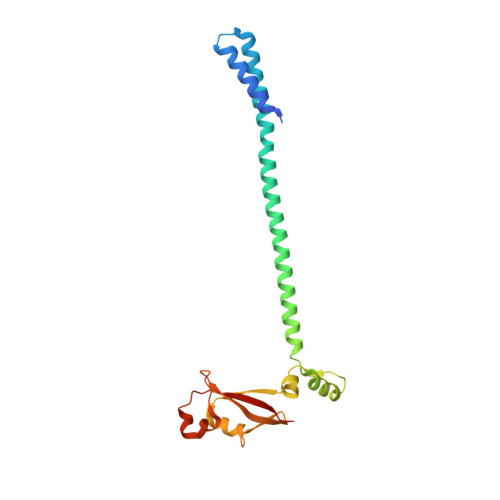
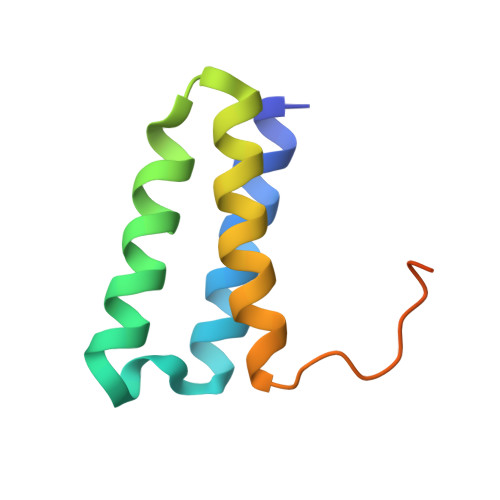
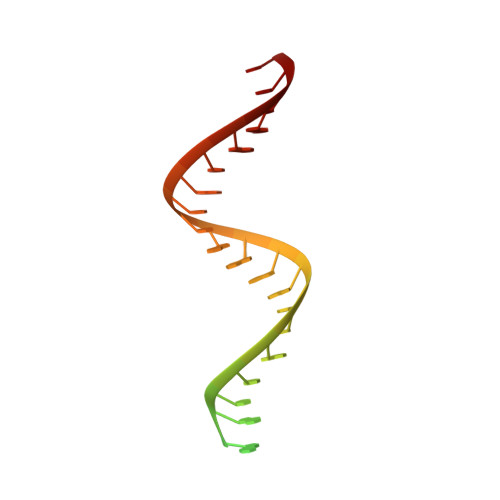
Coronaviruses are a diverse subfamily of viruses containing pathogens of humans and animals. This subfamily of viruses replicates their RNA genomes using a core polymerase complex composed of viral non-structural proteins: nsp7, nsp8 and nsp12. Most of our understanding of coronavirus molecular biology comes from the betacoronaviruses like SARS-CoV and SARS-CoV-2, the latter of which is the causative agent of COVID-19. In contrast, members of the alphacoronavirus genus are relatively understudied despite their importance in human and animal health. Here we have used cryo-electron microscopy to determine the structure of the alphacoronavirus porcine epidemic diarrhea virus (PEDV) core polymerase complex bound to RNA. Our structure shows an unexpected nsp8 stoichiometry in comparison to other published coronavirus polymerase structures. Biochemical analysis shows that the N-terminal extension of one nsp8 is not required for in vitro RNA synthesis for alpha and betacoronaviruses as previously hypothesized. Our work shows the importance of studying diverse coronaviruses to reveal aspects of coronavirus replication while also identifying areas of conservation to be targeted by antiviral drugs.
- Biochemistry Department, University of Wisconsin-Madison, Madison, WI 53706.
Organizational Affiliation: