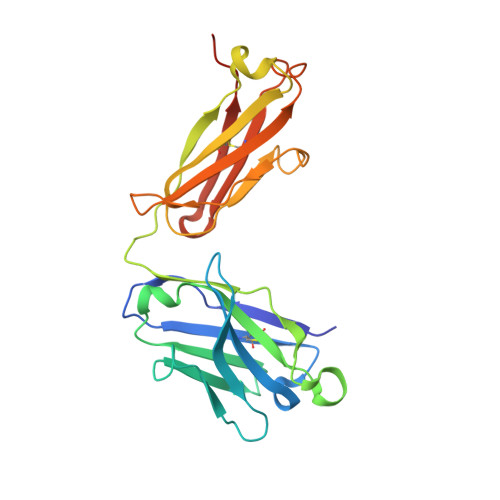
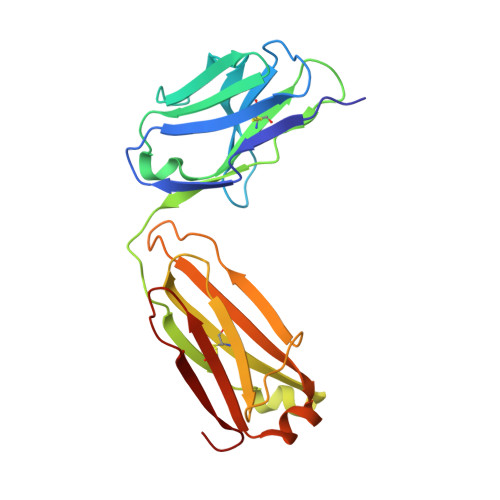
Structural basis for promiscuity in ligand recognition by yjdF riboswitch.
Krochmal, D., Roman, C., Lewicka, A., Shao, Y., Piccirilli, J.A.(2024) Cell Discov 10: 37-37
- PubMed: 38565535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-024-00663-2
- Primary Citation Related Structures:
8UIW, 8UTA - Department of Biochemistry and Molecular Biology, University of Chicago, Chicago, IL, USA.
Organizational Affiliation: