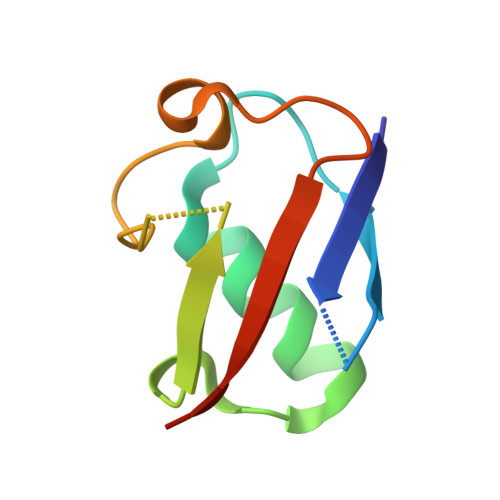
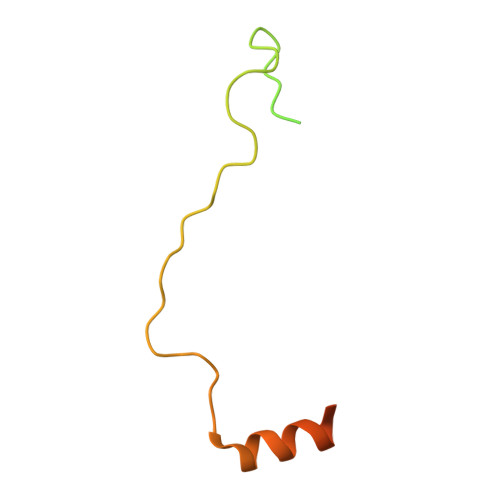
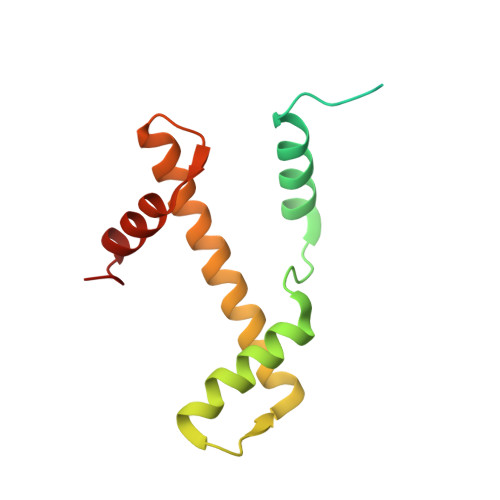
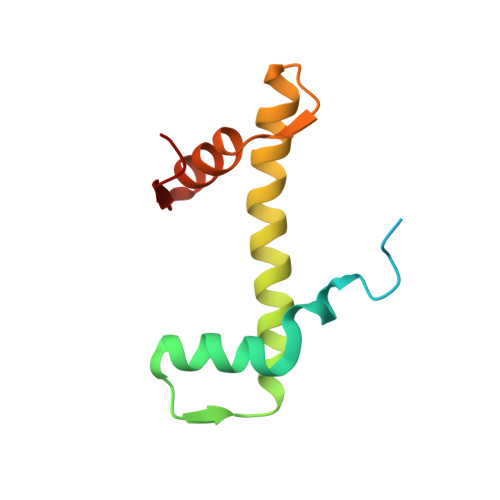
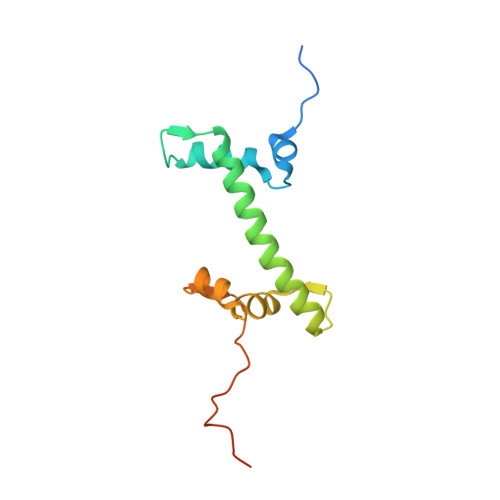
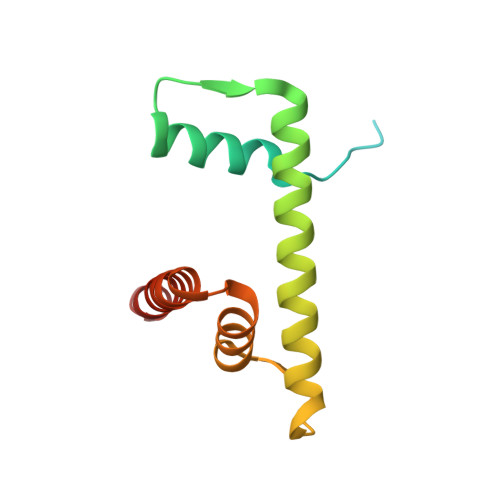
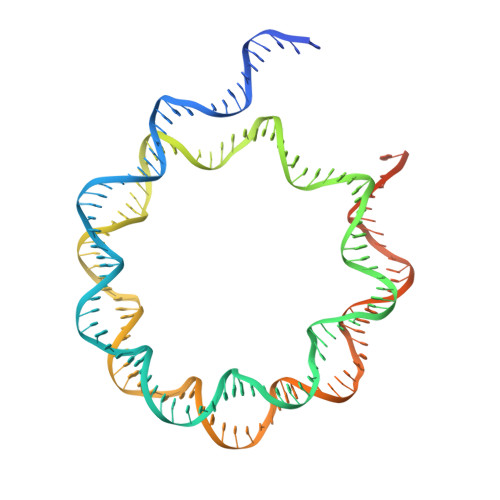
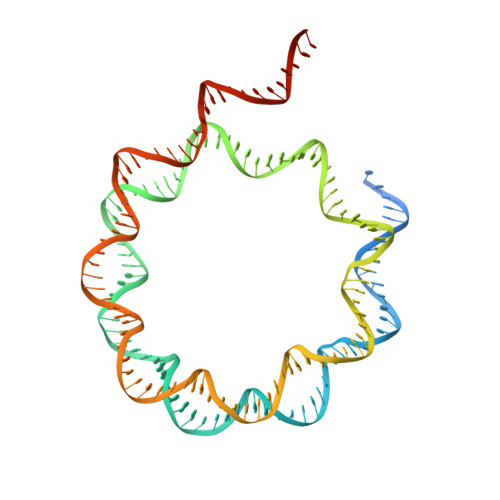
Structural basis for the H2AK119ub1-specific DNMT3A-nucleosome interaction.
Chen, X., Guo, Y., Zhao, T., Lu, J., Fang, J., Wang, Y., Wang, G.G., Song, J.(2024) Nat Commun 15: 6217-6217
- PubMed: 39043678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-50526-3
- Primary Citation Related Structures:
8U5H - PubMed Abstract:
Isoform 1 of DNA methyltransferase DNMT3A (DNMT3A1) specifically recognizes nucleosome monoubiquitylated at histone H2A lysine-119 (H2AK119ub1) for establishment of DNA methylation. Mis-regulation of this process may cause aberrant DNA methylation and pathogenesis. However, the molecular basis underlying DNMT3A1-nucleosome interaction remains elusive. Here we report the cryo-EM structure of DNMT3A1's ubiquitin-dependent recruitment (UDR) fragment complexed with H2AK119ub1-modified nucleosome. DNMT3A1 UDR occupies an extensive nucleosome surface, involving the H2A-H2B acidic patch, a surface groove formed by H2A and H3, nucleosomal DNA, and H2AK119ub1. The DNMT3A1 UDR's interaction with H2AK119ub1 affects the functionality of DNMT3A1 in cells in a context-dependent manner. Our structural and biochemical analysis also reveals competition between DNMT3A1 and JARID2, a cofactor of polycomb repression complex 2 (PRC2), for nucleosome binding, suggesting the interplay between different epigenetic pathways. Together, this study reports a molecular basis for H2AK119ub1-dependent DNMT3A1-nucleosome association, with important implications in DNMT3A1-mediated DNA methylation in development.
- Department of Biochemistry, University of California, Riverside, CA, 92521, USA.
Organizational Affiliation: