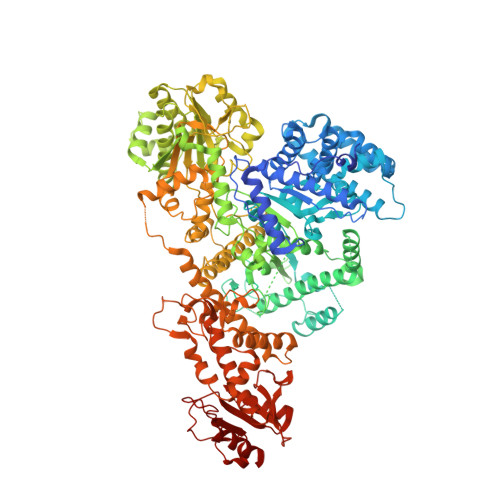
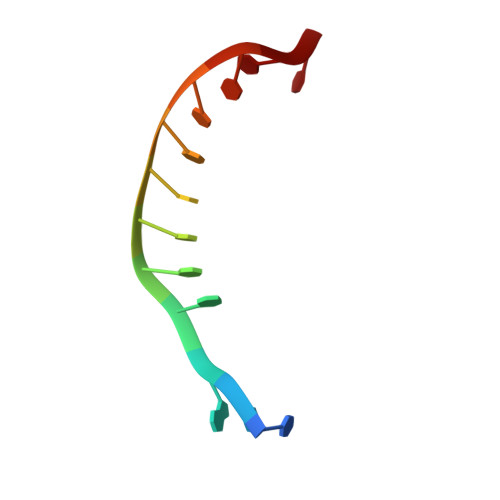
Plasmid targeting and destruction by the DdmDE bacterial defence system.
Bravo, J.P.K., Ramos, D.A., Fregoso Ocampo, R., Ingram, C., Taylor, D.W.(2024) Nature 630: 961-967
- PubMed: 38740055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07515-9
- Primary Citation Related Structures:
8U0J, 8U0U, 8U0W, 8U3K, 9BQV - PubMed Abstract:
Although eukaryotic Argonautes have a pivotal role in post-transcriptional gene regulation through nucleic acid cleavage, some short prokaryotic Argonaute variants (pAgos) rely on auxiliary nuclease factors for efficient foreign DNA degradation 1 . Here we reveal the activation pathway of the DNA defence module DdmDE system, which rapidly eliminates small, multicopy plasmids from the Vibrio cholerae seventh pandemic strain (7PET) 2 . Through a combination of cryo-electron microscopy, biochemistry and in vivo plasmid clearance assays, we demonstrate that DdmE is a catalytically inactive, DNA-guided, DNA-targeting pAgo with a distinctive insertion domain. We observe that the helicase-nuclease DdmD transitions from an autoinhibited, dimeric complex to a monomeric state upon loading of single-stranded DNA targets. Furthermore, the complete structure of the DdmDE-guide-target handover complex provides a comprehensive view into how DNA recognition triggers processive plasmid destruction. Our work establishes a mechanistic foundation for how pAgos utilize ancillary factors to achieve plasmid clearance, and provides insights into anti-plasmid immunity in bacteria.
- Department of Molecular Biosciences, University of Texas at Austin, Austin, TX, USA. jack.bravo@ist.ac.at.
Organizational Affiliation: