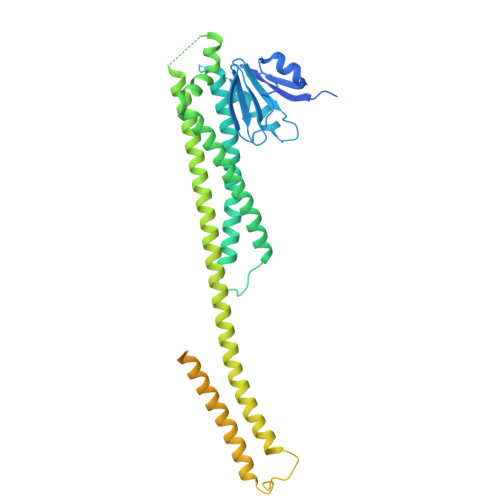
Structure and function of the human mitochondrial MRS2 channel.
He, Z., Tu, Y.C., Tsai, C.W., Mount, J., Zhang, J., Tsai, M.F., Yuan, P.(2025) Nat Struct Mol Biol 32: 459-468
- PubMed: 39609651 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01420-5
- Primary Citation Related Structures:
8TS1, 8TS2, 8TS3 - PubMed Abstract:
The human mitochondrial RNA splicing 2 protein (MRS2) has been implicated in Mg 2+ transport across mitochondrial inner membranes, thus having an important role in Mg 2+ homeostasis critical for mitochondrial integrity and function. However, the molecular mechanisms underlying its fundamental channel properties such as ion selectivity and regulation remain unclear. Here we present a structural and functional investigation of MRS2. Cryo-electron microscopy structures in various ionic conditions reveal a pentameric channel architecture and the molecular basis of ion permeation and potential regulation mechanisms. Electrophysiological analyses demonstrate that MRS2 is a Ca 2+ -regulated, nonselective channel permeable to Mg 2+ , Ca 2+ , Na + and K + , which contrasts with its prokaryotic ortholog, CorA, operating as a Mg 2+ -gated Mg 2+ channel. Moreover, a conserved arginine ring within the pore of MRS2 functions to restrict cation movements, thus preventing the channel from collapsing the proton motive force that drives mitochondrial adenosine triphosphate synthesis. Together, our results provide a molecular framework for further understanding MRS2 in mitochondrial function and disease.
- Department of Cell Biology and Physiology, Washington University School of Medicine, Saint Louis, MO, USA.
Organizational Affiliation: