Ultrapotent Broadly Neutralizing Human-llama Bispecific Antibodies against HIV-1.
Xu, J., Zhou, T., McKee, K., Zhang, B., Liu, C., Nazzari, A.F., Pegu, A., Shen, C.H., Becker, J.E., Bender, M.F., Chan, P., Changela, A., Chaudhary, R., Chen, X., Einav, T., Kwon, Y.D., Lin, B.C., Louder, M.K., Merriam, J.S., Morano, N.C., O'Dell, S., Olia, A.S., Rawi, R., Roark, R.S., Stephens, T., Teng, I.T., Tourtellott-Fogt, E., Wang, S., Yang, E.S., Shapiro, L., Tsybovsky, Y., Doria-Rose, N.A., Casellas, R., Kwong, P.D.(2024) Adv Sci (Weinh) 11: e2309268-e2309268
- PubMed: 38704686 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/advs.202309268
- Primary Citation Related Structures:
8TNG, 8TNH, 8TNI - PubMed Abstract:
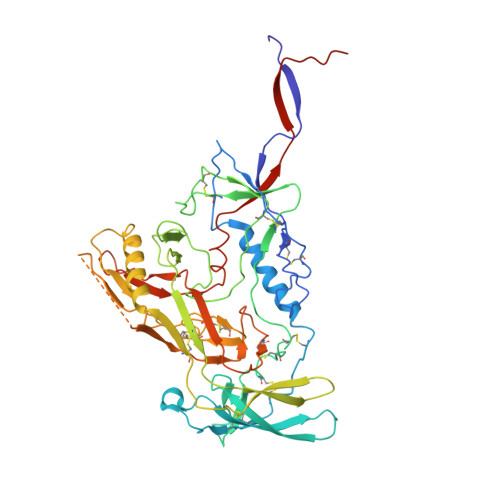
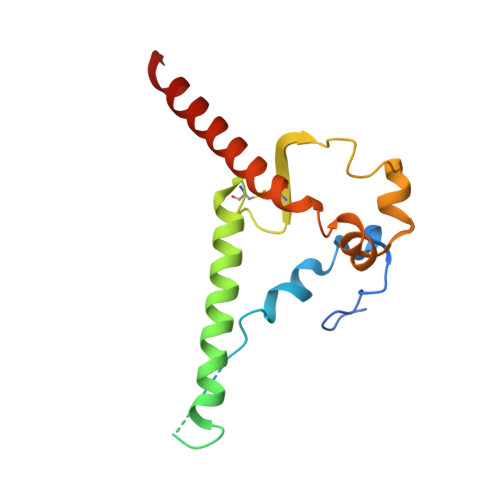
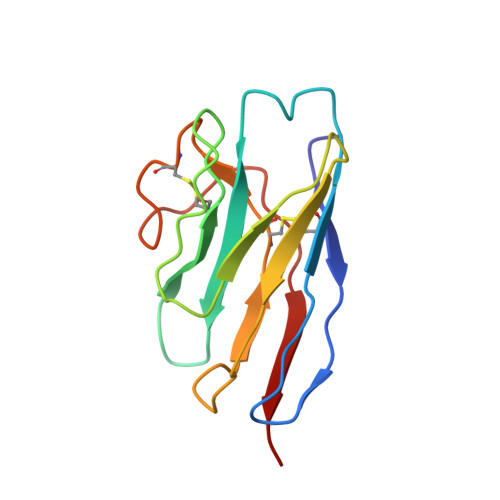
Broadly neutralizing antibodies are proposed as therapeutic and prophylactic agents against HIV-1, but their potency and breadth are less than optimal. This study describes the immunization of a llama with the prefusion-stabilized HIV-1 envelope (Env) trimer, BG505 DS-SOSIP, and the identification and improvement of potent neutralizing nanobodies recognizing the CD4-binding site (CD4bs) of vulnerability. Two of the vaccine-elicited CD4bs-targeting nanobodies, G36 and R27, when engineered into a triple tandem format with llama IgG2a-hinge region and human IgG1-constant region (G36×3-IgG2a and R27×3-IgG2a), neutralized 96% of a multiclade 208-strain panel at geometric mean IC 80 s of 0.314 and 0.033 µg mL -1 , respectively. Cryo-EM structures of these nanobodies in complex with Env trimer revealed the two nanobodies to neutralize HIV-1 by mimicking the recognition of the CD4 receptor. To enhance their neutralizing potency and breadth, nanobodies are linked to the light chain of the V2-apex-targeting broadly neutralizing antibody, CAP256V2LS. The resultant human-llama bispecific antibody CAP256L-R27×3LS exhibited ultrapotent neutralization and breadth exceeding other published HIV-1 broadly neutralizing antibodies, with pharmacokinetics determined in FcRn-Fc mice similar to the parent CAP256V2LS. Vaccine-elicited llama nanobodies, when combined with V2-apex broadly neutralizing antibodies, may therefore be able to fulfill anti-HIV-1 therapeutic and prophylactic clinical goals.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, 20892, USA.
Organizational Affiliation: