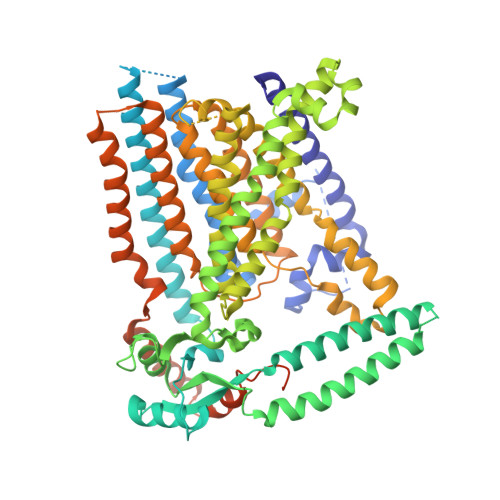
Structure of mechanically activated ion channel OSCA2.3 reveals mobile elements in the transmembrane domain.
Jojoa-Cruz, S., Burendei, B., Lee, W.H., Ward, A.B.(2024) Structure 32: 157
- PubMed: 38103547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2023.11.009
- Primary Citation Related Structures:
8T56, 8T57 - PubMed Abstract:
Members of the OSCA/TMEM63 family are mechanically activated ion channels and structures of some OSCA members have revealed the architecture of these channels and structural features that are potentially involved in mechanosensation. However, these structures are all in a similar state and information about the motion of different elements of the structure is limited, preventing a deeper understanding of how these channels work. Here, we used cryoelectron microscopy to determine high-resolution structures of Arabidopsis thaliana OSCA1.2 and OSCA2.3 in peptidiscs. The structure of OSCA1.2 matches previous structures of the same protein in different environments. Yet, in OSCA2.3, the TM6a-TM7 linker adopts a different conformation that constricts the pore on its cytoplasmic side. Furthermore, coevolutionary sequence analysis uncovered a conserved interaction between the TM6a-TM7 linker and the beam-like domain (BLD). Our results reveal conformational heterogeneity and differences in conserved interactions between the TMD and BLD among members of the OSCA family.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: