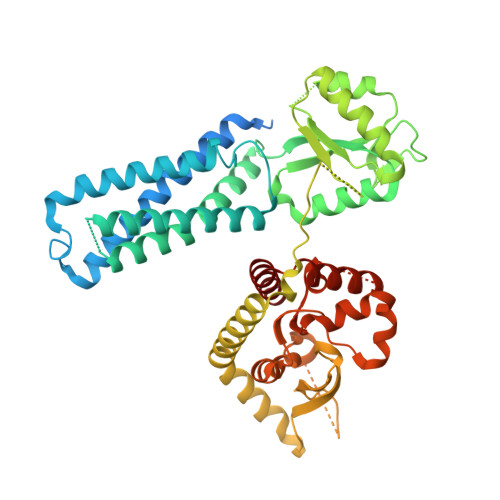
Mapping the architecture of the initiating phosphoglycosyl transferase from S. enterica O-antigen biosynthesis in a liponanoparticle.
Dodge, G.J., Anderson, A.J., He, Y., Liu, W., Viner, R., Imperiali, B.(2024) Elife 12
- PubMed: 38358918 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.91125
- Primary Citation Related Structures:
8T53 - PubMed Abstract:
Bacterial cell surface glycoconjugates are critical for cell survival and for interactions between bacteria and their hosts. Consequently, the pathways responsible for their biosynthesis have untapped potential as therapeutic targets. The localization of many glycoconjugate biosynthesis enzymes to the membrane represents a significant challenge for expressing, purifying, and characterizing these enzymes. Here, we leverage cutting-edge detergent-free methods to stabilize, purify, and structurally characterize WbaP, a phosphoglycosyl transferase (PGT) from the Salmonella enterica (LT2) O-antigen biosynthesis. From a functional perspective, these studies establish WbaP as a homodimer, reveal the structural elements responsible for dimerization, shed light on the regulatory role of a domain of unknown function embedded within WbaP, and identify conserved structural motifs between PGTs and functionally unrelated UDP-sugar dehydratases. From a technological perspective, the strategy developed here is generalizable and provides a toolkit for studying other classes of small membrane proteins embedded in liponanoparticles beyond PGTs.
- Department of Biology and Department of Chemistry, Massachusetts Institute of Technology, Cambridge, United States.
Organizational Affiliation: