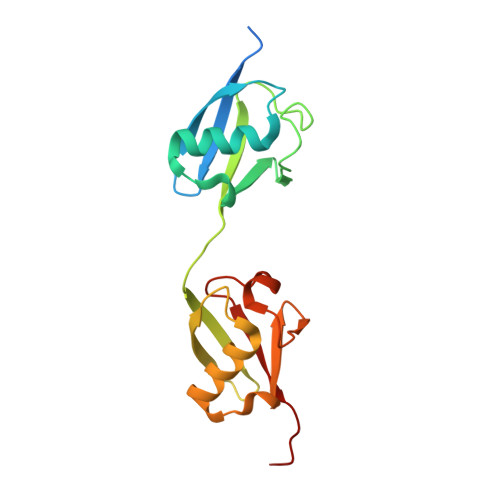
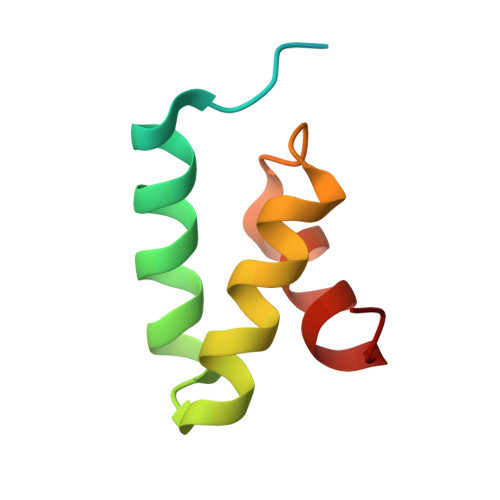
N4BP1 coordinates ubiquitin-dependent crosstalk within the I kappa B kinase family to limit Toll-like receptor signaling and inflammation.
Gitlin, A.D., Maltzman, A., Kanno, Y., Heger, K., Reja, R., Schubert, A.F., Wierciszewski, L.J., Pantua, H., Kapadia, S.B., Harris, S.F., Webster, J.D., Newton, K., Dixit, V.M.(2024) Immunity 57: 973
- PubMed: 38697117
- DOI: https://doi.org/10.1016/j.immuni.2024.04.004
- Primary Citation Related Structures:
8T48 - PubMed Abstract:
The ubiquitin-binding endoribonuclease N4BP1 potently suppresses cytokine production by Toll-like receptors (TLRs) that signal through the adaptor MyD88 but is inactivated via caspase-8-mediated cleavage downstream of death receptors, TLR3, or TLR4. Here, we examined the mechanism whereby N4BP1 limits inflammatory responses. In macrophages, deletion of N4BP1 prolonged activation of inflammatory gene transcription at late time points after TRIF-independent TLR activation. Optimal suppression of inflammatory cytokines by N4BP1 depended on its ability to bind polyubiquitin chains, as macrophages and mice-bearing inactivating mutations in a ubiquitin-binding motif in N4BP1 displayed increased TLR-induced cytokine production. Deletion of the noncanonical IκB kinases (ncIKKs), Tbk1 and Ikke, or their adaptor Tank phenocopied N4bp1 deficiency and enhanced macrophage responses to TLR1/2, TLR7, or TLR9 stimulation. Mechanistically, N4BP1 acted in concert with the ncIKKs to limit the duration of canonical IκB kinase (IKKα/β) signaling. Thus, N4BP1 and the ncIKKs serve as an important checkpoint against over-exuberant innate immune responses.
- Immunology Program, Sloan Kettering Institute, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA; Department of Pathology and Laboratory Medicine, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA; Immunology and Microbial Pathogenesis Program, Weill Cornell Graduate School of Medical Sciences, New York, NY 10065, USA. Electronic address: gitlina@mskcc.org.
Organizational Affiliation: