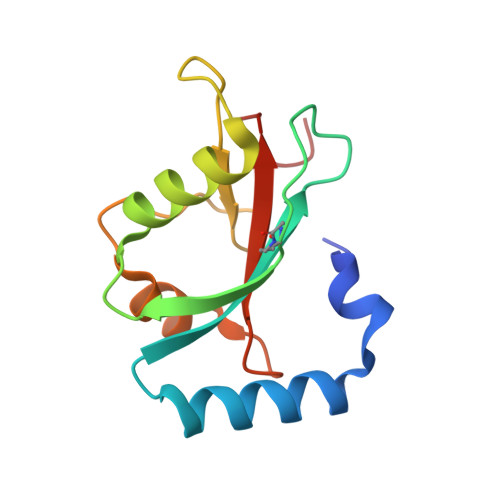
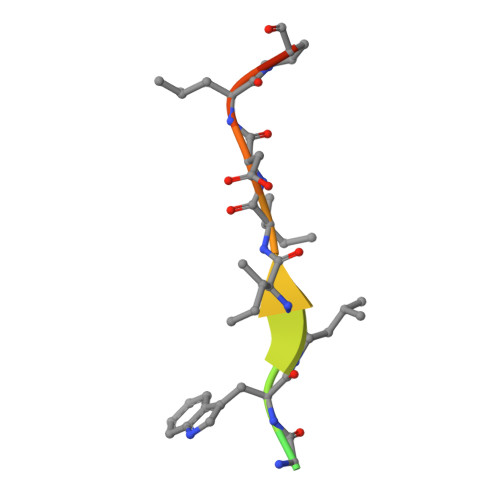
Structural and functional characterization of the role of acetylation on the interactions of the human Atg8-family proteins with the autophagy receptor TP53INP2/DOR.
Ali, M.G., Wahba, H.M., Igelmann, S., Cyr, N., Ferbeyre, G., Omichinski, J.G.(2024) Autophagy 20: 1948-1967
- PubMed: 38726830 Search on PubMed
- DOI: https://doi.org/10.1080/15548627.2024.2353443
- Primary Citation Related Structures:
8T31, 8T32, 8T33, 8T35, 8T36, 8T4T - PubMed Abstract:
The Atg8-family proteins (MAP1LC3/LC3A, LC3B, LC3C, GABARAP, GABARAPL1 and GABARAPL2) play a pivotal role in macroautophagy/autophagy through their ability to help form autophagosomes. Although autophagosomes form in the cytoplasm, nuclear levels of the Atg8-family proteins are significant. Recently, the nuclear/cytoplasmic shuttling of LC3B was shown to require deacetylation of two Lys residues (K49 and K51 in LC3B), which are conserved in Atg8-family proteins. To exit the nucleus, deacetylated LC3B must bind TP53INP2/TP53INP2 (tumor protein p53 inducible nuclear protein 2) through interaction with the LC3-interacting region (LIR) of TP53INP2 (TP53INP2LIR). To examine their selectivity for TP53INP2 and the role of the conserved Lys residues in Atg8-family proteins, we prepared the six human Atg8-family proteins and acetylated variants of LC3A and GABARAP for biophysical and structural characterization of their interactions with the TP53INP2LIR. Isothermal titration calorimetry (ITC) experiments demonstrate that this LIR binds preferentially to GABARAP subfamily proteins, and that only acetylation of the second Lys residue reduces binding to GABARAP and LC3A. Crystal structures of complexes with GABARAP and LC3A (acetylated and deacetylated) define a β-sheet in the TP53INP2LIR that determines the GABARAP selectivity and establishes the importance of acetylation at the second Lys. The in vitro results were confirmed in cells using acetyl-mimetic variants of GABARAP and LC3A to examine nuclear/cytoplasmic shuttling and colocalization with TP53INP2. Together, the results demonstrate that TP53INP2 shows selectivity to the GABARAP subfamily and acetylation at the second Lys of GABARAP and LC3A disrupts key interactions with TP53INP2 required for their nuclear/cytoplasmic shuttling.
- Département de Biochimie et Médicine Moléculaire, Université de Montréal, Montréal, QC, Canada.
Organizational Affiliation: