Biophysical characterization of PVR family interactions and therapeutic antibody recognition to TIGIT.
Diong, S.J., Jashnani, A., Drake, A.W., Bee, C., Findeisen, F., Dollinger, G., Wang, F., Rajpal, A., Strop, P., Lee, P.S.(2023) MAbs 15: 2253788-2253788
- PubMed: 37675979 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2023.2253788
- Primary Citation Related Structures:
8SZY - PubMed Abstract:
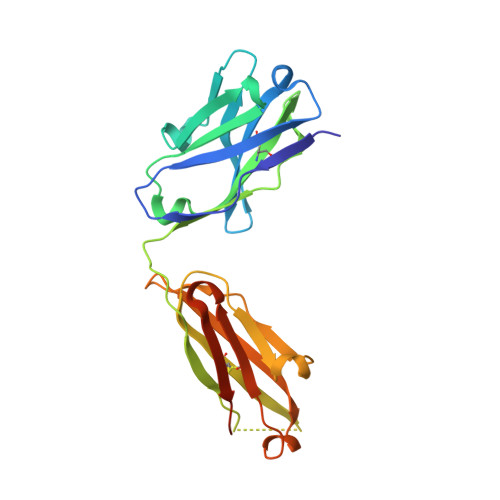
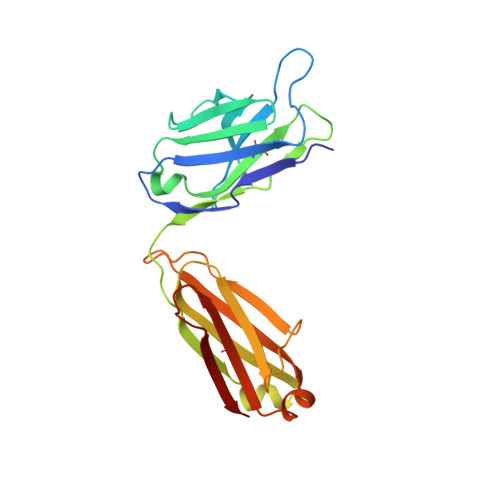
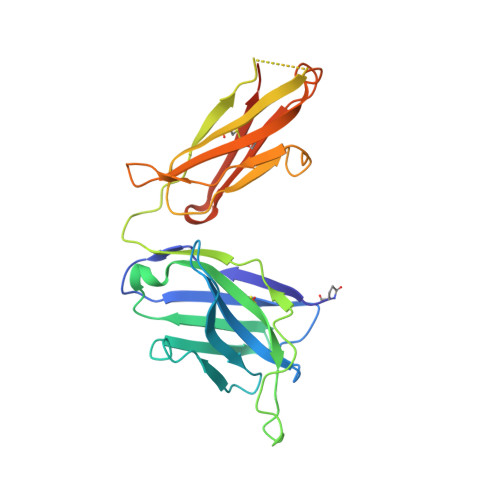
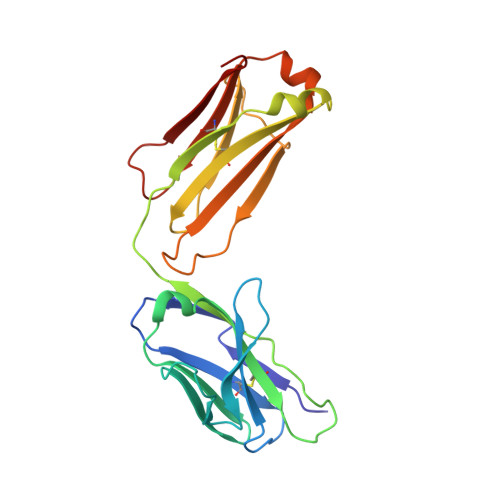
The clinical successes of immune checkpoint blockade have invigorated efforts to activate T cell-mediated responses against cancer. Targeting members of the PVR family, consisting of inhibitory receptors TIGIT, CD96, and CD112R, has been an active area of clinical investigation. In this study, the binding interactions and molecular assemblies of the PVR family receptors and ligands have been assessed in vitro . Furthermore, the anti-TIGIT monoclonal antibody BMS-986207 crystal structure in complex with TIGIT was determined and shows that the antibody binds an epitope that is commonly targeted by the CD155 ligand as well as other clinical anti-TIGIT antibodies. In contrast to previously proposed models, where TIGIT outcompetes costimulatory receptor CD226 for binding to CD155 due to much higher affinity (nanomolar range), our data rather suggest that PVR family members all engage in interactions with relatively weak affinity (micromolar range), including TIGIT and CD155 interactions. Thus, TIGIT and other PVR inhibitory receptors likely elicit immune suppression via increased surface expression rather than inherent differences in affinity. This work provides an improved foundational understanding of the PVR family network and mechanistic insight into therapeutic antibody intervention.
- Discovery Biologics, Bristol Myers Squibb, Redwood City, CA, USA.
Organizational Affiliation: