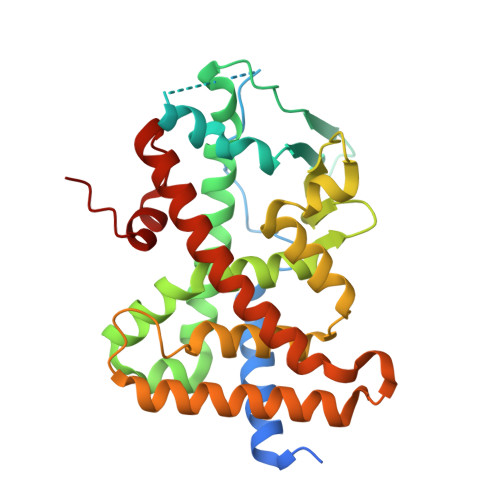
Ligand flexibility and binding pocket malleability cooperate to allow selective PXR activation by analogs of a promiscuous nuclear receptor ligand.
Huber, A.D., Poudel, S., Li, Y., Lin, W., Wu, J., Miller, D.J., Chen, T.(2023) Structure 31: 1545
- PubMed: 37729916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2023.08.020
- Primary Citation Related Structures:
8SZV - PubMed Abstract:
The human nuclear receptor (NR) family of transcription factors contains 48 proteins that bind lipophilic molecules. Approved NR therapies have had immense success treating various diseases, but lack of selectivity has hindered efforts to therapeutically target the majority of NRs due to unpredictable off-target effects. The synthetic ligand T0901317 was originally discovered as a potent agonist of liver X receptors (LXRα/β) but subsequently found to target additional NRs, with activation of pregnane X receptor (PXR) being as potent as that of LXRs. We previously showed that directed rigidity reduces PXR binding by T0901317 derivatives through unfavorable protein remodeling. Here, we use a similar approach to achieve selectivity for PXR over other T0901317-targeted NRs. One molecule, SJPYT-318, accomplishes selectivity by favorably utilizing PXR's flexible binding pocket and surprisingly binding in a new mode distinct from the parental T0901317. Our work provides a structure-guided framework to achieve NR selectivity from promiscuous compounds.
- Department of Chemical Biology and Therapeutics, St. Jude Children's Research Hospital, Memphis, TN 38105, USA.
Organizational Affiliation: