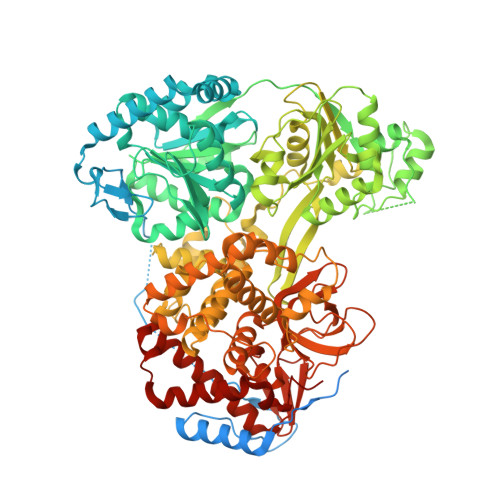
Development of assays to support identification and characterization of modulators of DExH-box helicase DHX9.
Gotur, D., Case, A., Liu, J., Sickmier, E.A., Holt, N., Knockenhauer, K.E., Yao, S., Lee, Y.T., Copeland, R.A., Buker, S.M., Boriack-Sjodin, P.A.(2023) SLAS Discov 28: 376-384
- PubMed: 37625785 Search on PubMed
- DOI: https://doi.org/10.1016/j.slasd.2023.08.006
- Primary Citation Related Structures:
8SZS - PubMed Abstract:
DHX9 is a DExH-box RNA helicase that utilizes hydrolysis of all four nucleotide triphosphates (NTPs) to power cycles of 3' to 5' directional movement to resolve and/or unwind double stranded RNA, DNA, and RNA/DNA hybrids, R-loops, triplex-DNA and G-quadraplexes. DHX9 activity is important for both viral amplification and maintaining genomic stability in cancer cells; therefore, it is a therapeutic target of interest for drug discovery efforts. Biochemical assays measuring ATP hydrolysis and oligonucleotide unwinding for DHX9 have been developed and characterized, and these assays can support high-throughput compound screening efforts under balanced conditions. Assay development efforts revealed DHX9 can use double stranded RNA with 18-mer poly(U) 3' overhangs and as well as significantly shorter overhangs at the 5' or 3' end as substrates. The enzymatic assays are augmented by a robust SPR assay for compound validation. A mechanism-derived inhibitor, GTPγS, was characterized as part of the validation of these assays and a crystal structure of GDP bound to cat DHX9 has been solved. In addition to enabling drug discovery efforts for DHX9, these assays may be extrapolated to other RNA helicases providing a valuable toolkit for this important target class.
- Accent Therapeutics, 1050 Waltham Street, Lexington, MA 02421, USA.
Organizational Affiliation: