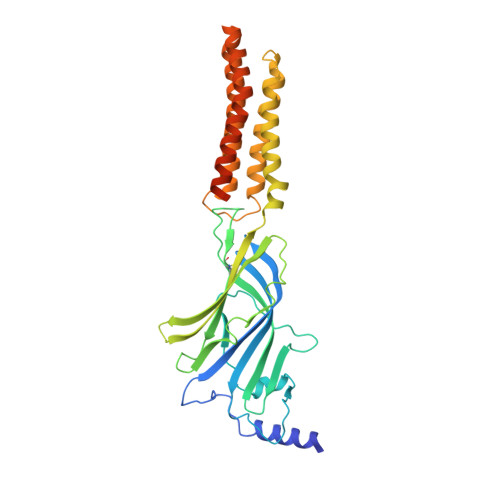
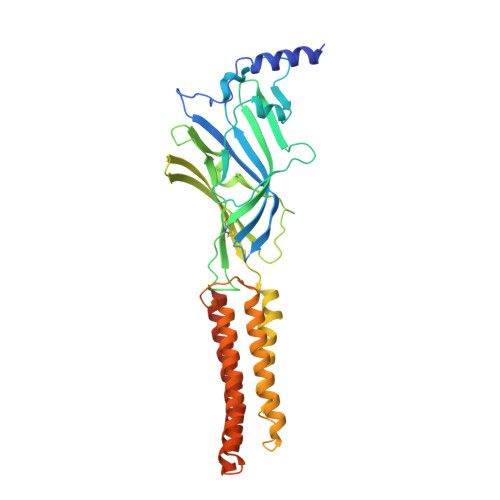
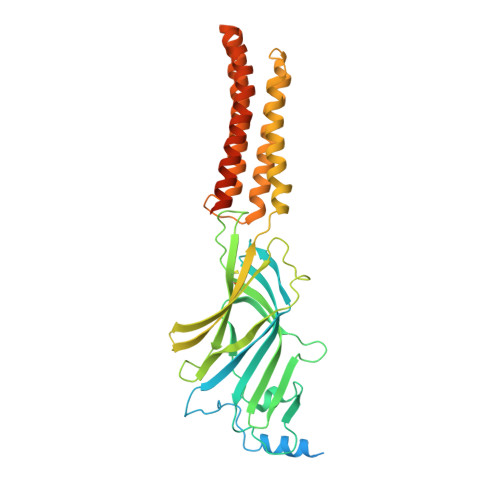
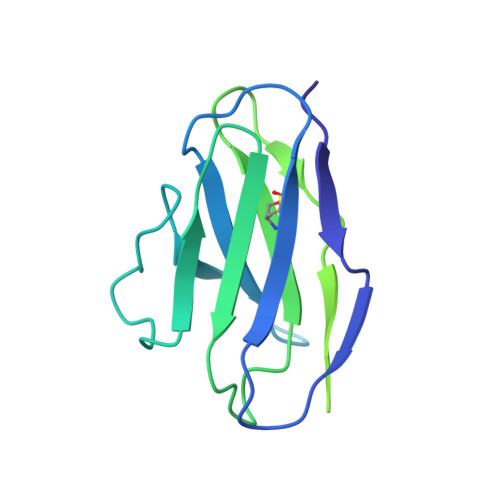
Structural insights into opposing actions of neurosteroids on GABA A receptors.
Legesse, D.H., Fan, C., Teng, J., Zhuang, Y., Howard, R.J., Noviello, C.M., Lindahl, E., Hibbs, R.E.(2023) Nat Commun 14: 5091-5091
- PubMed: 37607940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40800-1
- Primary Citation Related Structures:
8SGO, 8SI9, 8SID - PubMed Abstract:
γ-Aminobutyric acid type A (GABA A ) receptors mediate fast inhibitory signaling in the brain and are targets of numerous drugs and endogenous neurosteroids. A subset of neurosteroids are GABA A receptor positive allosteric modulators; one of these, allopregnanolone, is the only drug approved specifically for treating postpartum depression. There is a consensus emerging from structural, physiological and photolabeling studies as to where positive modulators bind, but how they potentiate GABA activation remains unclear. Other neurosteroids are negative modulators of GABA A receptors, but their binding sites remain debated. Here we present structures of a synaptic GABA A receptor bound to allopregnanolone and two inhibitory sulfated neurosteroids. Allopregnanolone binds at the receptor-bilayer interface, in the consensus potentiator site. In contrast, inhibitory neurosteroids bind in the pore. MD simulations and electrophysiology support a mechanism by which allopregnanolone potentiates channel activity and suggest the dominant mechanism for sulfated neurosteroid inhibition is through pore block.
- Department of Neuroscience, UT Southwestern Medical Center, Dallas, TX, USA.
Organizational Affiliation: